DNA sequences within glioma-derived extracellular vesicles can cross the intact blood-brain barrier and be detected in peripheral blood of patients
- PMID: 27902458
- PMCID: PMC5352065
- DOI: 10.18632/oncotarget.13635
DNA sequences within glioma-derived extracellular vesicles can cross the intact blood-brain barrier and be detected in peripheral blood of patients
Abstract
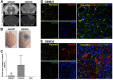
Tumor-cell-secreted extracellular vesicles (EVs) can cross the disrupted blood-brain barrier (BBB) into the bloodstream. However, in certain gliomas, the BBB remains intact, which might limit EVs release. To evaluate the ability of tumor-derived EVs to cross the BBB, we used an orthotopic xenotransplant mouse model of human glioma-cancer stem cells featuring an intact BBB. We demonstrated that all types of tumor cells-derived EVs-apoptotic bodies, shedding microvesicles and exosomes-cross the intact BBB and can be detected in the peripheral blood, which provides a minimally invasive method for their detection compared to liquid biopsies obtained from cerebrospinal fluid (CSF). Furthermore, these EVs can be readily distinguished from total murine EVs, since they carry human-specific DNA sequences relevant for GBM biology. In a small cohort of glioma patients, we finally demonstrated that peripheral blood EVs cargo can be successfully used to detect the presence of IDH1G395A, an essential biomarker in the current management of human glioma.
Keywords: biomarkers; blood-brain barrier; brain tumors; extracellular vesicles.
Conflict of interest statement
N.G.R., J.C.N., C.B.I. and A.A.S. declare competing financial interest in this work related to a patent pending on the use of a method for the detection of gene mutations in DNA from extracellular vesicles. (Application number: 16382028.5 – 1403). The other authors have no conflict of interests to declare.
Figures



Similar articles
-
The emerging clinical potential of circulating extracellular vesicles for non-invasive glioma diagnosis and disease monitoring.Brain Tumor Pathol. 2019 Apr;36(2):29-39. doi: 10.1007/s10014-019-00335-0. Epub 2019 Mar 11. Brain Tumor Pathol. 2019. PMID: 30859343 Review.
-
Tumor-Derived Extracellular Vesicles Breach the Intact Blood-Brain Barrier via Transcytosis.ACS Nano. 2019 Dec 24;13(12):13853-13865. doi: 10.1021/acsnano.9b04397. Epub 2019 Sep 10. ACS Nano. 2019. PMID: 31479239 Free PMC article.
-
The Transport Mechanism of Extracellular Vesicles at the Blood-Brain Barrier.Curr Pharm Des. 2017;23(40):6206-6214. doi: 10.2174/1381612823666170913164738. Curr Pharm Des. 2017. PMID: 28914201 Review.
-
Canine glioblastoma-derived extracellular vesicles as precise carriers for glioblastoma imaging: Targeting across the blood-brain barrier.Biomed Pharmacother. 2024 Mar;172:116201. doi: 10.1016/j.biopha.2024.116201. Epub 2024 Feb 1. Biomed Pharmacother. 2024. PMID: 38306846
-
Targeting and Crossing the Blood-Brain Barrier with Extracellular Vesicles.Cells. 2020 Apr 1;9(4):851. doi: 10.3390/cells9040851. Cells. 2020. PMID: 32244730 Free PMC article. Review.
Cited by
-
3'-UTR Sequence of Exosomal NANOGP8 DNA as an Extracellular Vesicle-Localization Signal.Int J Mol Sci. 2024 Jul 2;25(13):7294. doi: 10.3390/ijms25137294. Int J Mol Sci. 2024. PMID: 39000405 Free PMC article.
-
The role and application of small extracellular vesicles in glioma.Cancer Cell Int. 2024 Jun 29;24(1):229. doi: 10.1186/s12935-024-03389-z. Cancer Cell Int. 2024. PMID: 38951882 Free PMC article. Review.
-
Exploratory study on microRNA profiles from plasma-derived extracellular vesicles in Alzheimer's disease and dementia with Lewy bodies.Transl Neurodegener. 2019 Oct 3;8:31. doi: 10.1186/s40035-019-0169-5. eCollection 2019. Transl Neurodegener. 2019. PMID: 31592314 Free PMC article.
-
New windows into the brain: Central nervous system-derived extracellular vesicles in blood.Prog Neurobiol. 2019 Apr;175:96-106. doi: 10.1016/j.pneurobio.2019.01.005. Epub 2019 Jan 25. Prog Neurobiol. 2019. PMID: 30685501 Free PMC article. Review.
-
Delivering the Promise of Gene Therapy with Nanomedicines in Treating Central Nervous System Diseases.Adv Sci (Weinh). 2022 Sep;9(26):e2201740. doi: 10.1002/advs.202201740. Epub 2022 Jul 18. Adv Sci (Weinh). 2022. PMID: 35851766 Free PMC article. Review.
References
-
- Chinot OL, Wick W, Mason W, Henriksson R, Saran F, Nishikawa R, Carpentier AF, Hoang-Xuan K, Kavan P, Cernea D, Brandes AA, Hilton M, Abrey L, et al. Bevacizumab plus radiotherapy-temozolomide for newly diagnosed glioblastoma. N Engl J Med. 2014;370:709–22. doi: 10.1056/NEJMoa1308345. - DOI - PubMed
-
- Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta Neuropathol. 2016;131:803–20. doi: 10.1007/s00401-016-1545-1. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

