Disease evolution and heterogeneity in bilateral breast cancer
- PMID: 27904775
- PMCID: PMC5126277
Disease evolution and heterogeneity in bilateral breast cancer
Abstract
Bilateral breast cancers (BBC) are currently treated as independent tumors arising in the same patient. Herein, we investigated whether BBC indeed evolve independently at the genomic level. We examined paired targeted next generation sequencing genotypes from 155 paraffin tumors corresponding to 76 BBC patients (75 women and one man; 52 concurrent and 24 metachronous), for coding mutations (amino acid changing, minor allele frequency <0.1%) and single nucleotide polymorphism (SNP) zygosity. Germline genotypes were available for 29 patients. Mutations were present in 80 tumors (54/76 patients; 71%), were mostly tumor-private (90%), more frequent in TP53 (19%), PIK3CA (14%), CDH1, GATA3, MLL3. TP53 mutations were more frequent in metachronous tumors (P<0.001); hormone receptor negative (P<0.001); with higher Ki-67 (P=0.002); and, in younger patients (P=0.01). Hypermutated tumors, all TP53 mutated, were diagnosed as the first incidence in 5 patients; their metachronous counterparts were mutation poor without TP53 involvement. Paired tumors shared common mutations at intratumoral frequency >20% in 10/54 comparable BBC (18.5%), 8/10 concurrent. SNP zygosity status was less preserved in metachronous, compared to concurrent disease. Pathogenic germline mutations were present in 10/29 patients, 9 in BRCA1 and one in TP53 (p.Phe341Val, first report in the germline). BBC demonstrated extensive inter- and intra-patient heterogeneity in the present thus far largest series of corresponding paired genotypes. The majority evolve independently and unpredictably, supporting current clinical practice. A considerable minority though, retains clonal origin and may be regarded as a distinct group for therapeutic interventions among concurrent BBC.
Keywords: Bilateral; breast cancer; clonality; coding mutations; concurrent; contralateral; hypermutation; metachronous; targeted next generation sequencing.
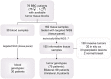
Figures




Similar articles
-
The fate of BRCA1-related germline mutations in triple-negative breast tumors.Am J Cancer Res. 2017 Jan 1;7(1):98-114. eCollection 2017. Am J Cancer Res. 2017. PMID: 28123851 Free PMC article.
-
Clinical characteristics and clinicopathological correlations of bilateral breast cancer in China: A multicenter study from Chinese Society of Breast Surgery (CSBrS-006).Chin J Cancer Res. 2021 Feb 28;33(1):27-32. doi: 10.21147/j.issn.1000-9604.2021.01.03. Chin J Cancer Res. 2021. PMID: 33707925 Free PMC article.
-
Addition of triple negativity of breast cancer as an indicator for germline mutations in predisposing genes increases sensitivity of clinical selection criteria.BMC Cancer. 2018 Sep 26;18(1):926. doi: 10.1186/s12885-018-4821-8. BMC Cancer. 2018. PMID: 30257646 Free PMC article.
-
Characterizations of Cancer Gene Mutations in Chinese Metastatic Breast Cancer Patients.Front Oncol. 2020 Jun 30;10:1023. doi: 10.3389/fonc.2020.01023. eCollection 2020. Front Oncol. 2020. PMID: 32695676 Free PMC article.
-
Li-Fraumeni syndrome: not a straightforward diagnosis anymore-the interpretation of pathogenic variants of low allele frequency and the differences between germline PVs, mosaicism, and clonal hematopoiesis.Breast Cancer Res. 2019 Sep 18;21(1):107. doi: 10.1186/s13058-019-1193-1. Breast Cancer Res. 2019. PMID: 31533767 Free PMC article. Review.
Cited by
-
A comparative clinicopathological and survival analysis of synchronous bilateral breast cancers.Histol Histopathol. 2022 Aug;37(8):791-802. doi: 10.14670/HH-18-449. Epub 2022 Mar 14. Histol Histopathol. 2022. PMID: 35285011
-
The prognostic comparison among unilateral, bilateral, synchronous bilateral, and metachronous bilateral breast cancer: A meta-analysis of studies from recent decade (2008-2018).Cancer Med. 2019 Jun;8(6):2908-2918. doi: 10.1002/cam4.2198. Epub 2019 Apr 30. Cancer Med. 2019. PMID: 31038845 Free PMC article.
-
Pathogenic mutations and overall survival in 3,084 patients with cancer: the Hellenic Cooperative Oncology Group Precision Medicine Initiative.Oncotarget. 2020 Jan 7;11(1):1-14. doi: 10.18632/oncotarget.27338. eCollection 2020 Jan 7. Oncotarget. 2020. PMID: 32002119 Free PMC article.
-
Frequency and diagnostic outcome of bilateral recall at screening mammography.Int J Cancer. 2021 Jan 1;148(1):48-56. doi: 10.1002/ijc.33187. Epub 2020 Jul 17. Int J Cancer. 2021. PMID: 32621785 Free PMC article.
-
The most likely but largely ignored triggering factor for breast (or all) cancer invasion.J Cancer. 2023 Feb 27;14(4):573-590. doi: 10.7150/jca.82291. eCollection 2023. J Cancer. 2023. PMID: 37057291 Free PMC article.
References
-
- SEER Database: Surveillance EaER, (SEER) Available from: http://seer.cancer.gov/
-
- Gong SJ, Rha SY, Jeung HC, Roh JK, Yang WI, Chung HC. Bilateral breast cancer: differential diagnosis using histological and biological parameters. Jpn J Clin Oncol. 2007;37:487–492. - PubMed
-
- Narod SA. Bilateral breast cancers. Nat Rev Clin Oncol. 2014;11:157–166. - PubMed
-
- Hartman M, Czene K, Reilly M, Adolfsson J, Bergh J, Adami HO, Dickman PW, Hall P. Incidence and prognosis of synchronous and metachronous bilateral breast cancer. J. Clin. Oncol. 2007;25:4210–4216. - PubMed
-
- Intra M, Rotmensz N, Viale G, Mariani L, Bonanni B, Mastropasqua MG, Galimberti V, Gennari R, Veronesi P, Colleoni M, Tousimis E, Galli A, Goldhirsch A, Veronesi U. Clinicopathologic characteristics of 143 patients with synchronous bilateral invasive breast carcinomas treated in a single institution. Cancer. 2004;101:905–912. - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
