Production of Phloroglucinol, a Platform Chemical, in Arabidopsis using a Bacterial Gene
- PMID: 27924918
- PMCID: PMC5141504
- DOI: 10.1038/srep38483
Production of Phloroglucinol, a Platform Chemical, in Arabidopsis using a Bacterial Gene
Abstract
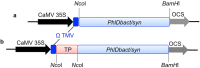
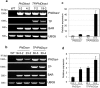
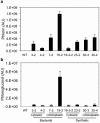
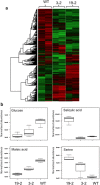
Phloroglucinol (1,3,5-trihydroxybenzene; PG) and its derivatives are phenolic compounds that are used for various industrial applications. Current methods to synthesize PG are not sustainable due to the requirement for carbon-based precursors and co-production of toxic byproducts. Here, we describe a more sustainable production of PG using plants expressing a native bacterial or a codon-optimized synthetic PhlD targeted to either the cytosol or chloroplasts. Transgenic lines were analyzed for the production of PG using gas and liquid chromatography coupled to mass spectroscopy. Phloroglucinol was produced in all transgenic lines and the line with the highest PhlD transcript level showed the most accumulation of PG. Over 80% of the produced PG was glycosylated to phlorin. Arabidopsis leaves have the machinery to glycosylate PG to form phlorin, which can be hydrolyzed enzymatically to produce PG. Furthermore, the metabolic profile of plants with PhlD in either the cytosol or chloroplasts was altered. Our results provide evidence that plants can be engineered to produce PG using a bacterial gene. Phytoproduction of PG using a bacterial gene paves the way for further genetic manipulations to enhance the level of PG with implications for the commercial production of this important platform chemical in plants.
Figures




References
-
- Xu J., Dolan M. C., Medrano G., Cramer C. L. & Weathers P. J. Green factory: Plants as bioproduction platforms for recombinant proteins. Biotechnology Advances 30, 1171–1184 (2012). - PubMed
-
- Abdel-Ghany S. E., Golovkin M. & Reddy A. S. In Plant Biology and Biotechnology Vol. II (eds Manchikatla Venkat Rajam Bir Bahadur, Sahijram Leela & Krishnamurthy K. V.) Ch. 28, 551–578 (Springer India, 2015).
-
- Agriculture U. S. D. o. US Biobases Products: Market Potential and projections through 2025. 1–294 (2008).
-
- Yoon J. M., Zhao L. & Shanks J. V. Metabolic Engineering with Plants for a Sustainable Biobased Economy. Annual Review of Chemical and Biomolecular Engineering 4, 211–237 (2013). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

