Reconstructing genetic history of Siberian and Northeastern European populations
- PMID: 27965293
- PMCID: PMC5204334
- DOI: 10.1101/gr.202945.115
Reconstructing genetic history of Siberian and Northeastern European populations
Abstract
Siberia and Northwestern Russia are home to over 40 culturally and linguistically diverse indigenous ethnic groups, yet genetic variation and histories of peoples from this region are largely uncharacterized. We present deep whole-genome sequencing data (∼38×) from 28 individuals belonging to 14 distinct indigenous populations from that region. We combined these data sets with additional 32 modern-day and 46 ancient human genomes to reconstruct genetic histories of several indigenous Northern Eurasian populations. We found that Siberian and East Asian populations shared 38% of their ancestry with a 45,000-yr-old Ust'-Ishim individual who was previously believed to have no modern-day descendants. Western Siberians trace 57% of their ancestry to ancient North Eurasians, represented by the 24,000-yr-old Siberian Mal'ta boy MA-1. Eastern Siberian populations formed a distinct sublineage that separated from other East Asian populations ∼10,000 yr ago. In addition, we uncovered admixtures between Siberians and Eastern European hunter-gatherers from Samara, Karelia, Hungary, and Sweden (from 8000-6600 yr ago); Yamnaya people (5300-4700 yr ago); and modern-day Northeastern Europeans. Our results provide new insights into genetic histories of Siberian and Northeastern European populations and evidence of ancient gene flow from Siberia into Europe.
© 2017 Wong et al.; Published by Cold Spring Harbor Laboratory Press.
Figures

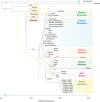


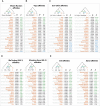


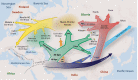
References
-
- Allentoft ME, Sikora M, Sjögren KG, Rasmussen S, Rasmussen M, Stenderup J, Damgaard PB, Schroeder H, Ahlström T, Vinner L, et al. 2015. Population genomics of Bronze Age Eurasia. Nature 522: 167–172. - PubMed
-
- Bunak VV. 1956. Human races and ways of their formation. Sov Ethnogr 1: 86–105 (in Russian).
-
- Bunak VV. 1965. Problems of the genesis of races. In The origin and ethnic history of Russian people (ed. Bunak VV), pp. 174–190. Nauka, Moscow (in Russian).
-
- Cheboksarov NN, Trofimova TA. 1941. Anthropological survey of Mansi. Short Reports of Institute of History of Material Culture 9: 28–36.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
