Within-Host Evolution of Simian Arteriviruses in Crab-Eating Macaques
- PMID: 27974564
- PMCID: PMC5286893
- DOI: 10.1128/JVI.02231-16
Within-Host Evolution of Simian Arteriviruses in Crab-Eating Macaques
Abstract
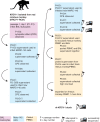
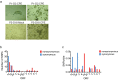
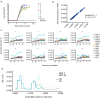
Simian arteriviruses are a diverse clade of viruses infecting captive and wild nonhuman primates. We recently reported that Kibale red colobus virus 1 (KRCV-1) causes a mild and self-limiting disease in experimentally infected crab-eating macaques, while simian hemorrhagic fever virus (SHFV) causes lethal viral hemorrhagic fever. Here we characterize how these viruses evolved during replication in cell culture and in experimentally infected macaques. During passage in cell culture, 68 substitutions that were localized in open reading frames (ORFs) likely associated with host cell entry and exit became fixed in the KRCV-1 genome. However, we did not detect any strong signatures of selection during replication in macaques. We uncovered patterns of evolution that were distinct from those observed in surveys of wild red colobus monkeys, suggesting that these species may exert different adaptive challenges for KRCV-1. During SHFV infection, we detected signatures of selection on ORF 5a and on a small subset of sites in the genome. Overall, our data suggest that patterns of evolution differ markedly among simian arteriviruses and among host species.
Importance: Certain RNA viruses can cross species barriers and cause disease in new hosts. Simian arteriviruses are a diverse group of related viruses that infect captive and wild nonhuman primates, with associated disease severity ranging from apparently asymptomatic infections to severe, viral hemorrhagic fevers. We infected nonhuman primate cell cultures and then crab-eating macaques with either simian hemorrhagic fever virus (SHFV) or Kibale red colobus virus 1 (KRCV-1) and assessed within-host viral evolution. We found that KRCV-1 quickly acquired a large number of substitutions in its genome during replication in cell culture but that evolution in macaques was limited. In contrast, we detected selection focused on SHFV ORFs 5a and 5, which encode putative membrane proteins. These patterns suggest that in addition to diverse pathogenic phenotypes, these viruses may also exhibit distinct patterns of within-host evolution both in vitro and in vivo.
Keywords: evolution; simian arterivirus; simian hemorrhagic fever virus; virology; within-host diversity.
Copyright © 2017 American Society for Microbiology.
Figures

References
-
- Allen AM, Palmer AE, Tauraso NM, Shelokov A. 1968. Simian hemorrhagic fever. II. Studies in pathology. Am J Trop Med Hyg 17:413–421. - PubMed
-
- Palmer AE, Allen AM, Tauraso NM, Shelokov A. 1968. Simian hemorrhagic fever. I. Clinical and epizootiologic aspects of an outbreak among quarantined monkeys. Am J Trop Med Hyg 17:404–412. - PubMed
-
- Tauraso NM, Shelokov A, Palmer AE, Allen AM. 1968. Simian hemorrhagic fever. 3. Isolation and characterization of a viral agent. Am J Trop Med Hyg 17:422–431. - PubMed
-
- Abildgaard C, Harrison J, Espana C, Spangler W, Gribble D. 1975. Simian hemorrhagic fever: studies of coagulation and pathology. Am J Trop Med Hyg 24:537–544. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

