The accommodation index measures the perturbation associated with insertions and deletions in coiled-coils: Application to understand signaling in histidine kinases
- PMID: 27977891
- PMCID: PMC5326573
- DOI: 10.1002/pro.3095
The accommodation index measures the perturbation associated with insertions and deletions in coiled-coils: Application to understand signaling in histidine kinases
Abstract
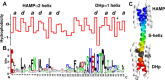
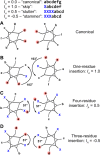
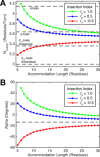
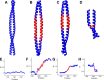
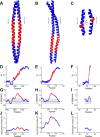
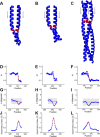
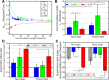
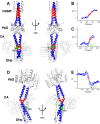
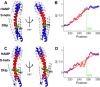
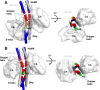
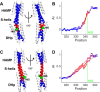
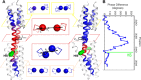
Coiled-coils are essential components of many protein complexes. First discovered in structural proteins such as keratins, they have since been found to figure largely in the assembly and dynamics required for diverse functions, including membrane fusion, signal transduction and motors. Coiled-coils have a characteristic repeating seven-residue geometric and sequence motif, which is sometimes interrupted by the insertion of one or more residues. Such insertions are often highly conserved and critical to interdomain communication in signaling proteins such as bacterial histidine kinases. Here we develop the "accommodation index" as a parameter that allows automatic detection and classification of insertions based on the three dimensional structure of a protein. This method allows precise identification of the type of insertion and the "accommodation length" over which the insertion is structurally accommodated. A simple theory is presented that predicts the structural perturbations of 1, 3, 4 residue insertions as a function of the length over which the insertion is accommodated. Analysis of experimental structures is in good agreement with theory, and shows that short accommodation lengths give rise to greater perturbation of helix packing angles, changes in local helical phase, and increased structural asymmetry relative to long accommodation lengths. Cytoplasmic domains of histidine kinases in different signaling states display large changes in their accommodation lengths, which can now be seen to underlie diverse structural transitions including symmetry/asymmetry and local variations in helical phase that accompany signal transduction.
Keywords: coiled-coil; heptad repeat; histidine kinase; protein design; protein structure analysis; theory.
© 2016 The Protein Society.
Figures

Similar articles
-
Dimer Asymmetry and Light Activation Mechanism in Brucella Blue-Light Sensor Histidine Kinase.mBio. 2021 Apr 20;12(2):e00264-21. doi: 10.1128/mBio.00264-21. mBio. 2021. PMID: 33879593 Free PMC article.
-
Signal Sensing and Transduction by Histidine Kinases as Unveiled through Studies on a Temperature Sensor.Acc Chem Res. 2017 Jun 20;50(6):1359-1366. doi: 10.1021/acs.accounts.6b00593. Epub 2017 May 5. Acc Chem Res. 2017. PMID: 28475313 Review.
-
Flexible Hinges in Bacterial Chemoreceptors.J Bacteriol. 2018 Feb 7;200(5):e00593-17. doi: 10.1128/JB.00593-17. Print 2018 Mar 1. J Bacteriol. 2018. PMID: 29229700 Free PMC article.
-
Transmembrane Prolines Mediate Signal Sensing and Decoding in Bacillus subtilis DesK Histidine Kinase.mBio. 2019 Nov 26;10(6):e02564-19. doi: 10.1128/mBio.02564-19. mBio. 2019. PMID: 31772055 Free PMC article.
-
Transmembrane Signal Transduction in Two-Component Systems: Piston, Scissoring, or Helical Rotation?Bioessays. 2018 Feb;40(2). doi: 10.1002/bies.201700197. Epub 2017 Dec 27. Bioessays. 2018. PMID: 29280502 Review.
Cited by
-
The coiled-coil domain of Escherichia coli FtsLB is a structurally detuned element critical for modulating its activation in bacterial cell division.J Biol Chem. 2022 Jan;298(1):101460. doi: 10.1016/j.jbc.2021.101460. Epub 2021 Dec 4. J Biol Chem. 2022. PMID: 34871549 Free PMC article.
-
Mechanism of conditional partner selectivity in MITF/TFE family transcription factors with a conserved coiled coil stammer motif.Nucleic Acids Res. 2020 Jan 24;48(2):934-948. doi: 10.1093/nar/gkz1104. Nucleic Acids Res. 2020. PMID: 31777941 Free PMC article.
-
Conformational Changes of an Interdomain Linker Mediate Mechanical Signal Transmission in Sensor Kinase BvgS.J Bacteriol. 2017 Aug 22;199(18):e00114-17. doi: 10.1128/JB.00114-17. Print 2017 Sep 15. J Bacteriol. 2017. PMID: 28507245 Free PMC article.
-
AlphaFold2 captures the conformational landscape of the HAMP signaling domain.Protein Sci. 2024 Jan;33(1):e4846. doi: 10.1002/pro.4846. Protein Sci. 2024. PMID: 38010737 Free PMC article.
-
High-resolution structure prediction of β-barrel membrane proteins.Proc Natl Acad Sci U S A. 2018 Feb 13;115(7):1511-1516. doi: 10.1073/pnas.1716817115. Epub 2018 Jan 29. Proc Natl Acad Sci U S A. 2018. PMID: 29378944 Free PMC article.
References
-
- Lupas A (1996) Coiled‐coils: new structures and new functions. Trends Biochem Sci 21:375–382. - PubMed
-
- Cohen C, Parry DAD (1990) a‐Helical coiled‐coils and bundles: how to design an a‐helical protein. Proteins 7:1–15. - PubMed
-
- Weissenhorn W, Dessen A, Calder LJ, Harrison SC, Skehel JJ, Wiley DC (1999) Structural basis for membrane fusion by enveloped viruses. Mol Membr Biol 16:3. - PubMed
-
- Dutch RE, Jardetzky TS, Lamb RA (2000) Virus membrane fusion proteins: biological machines that undergo a metamorphosis. Biosci Rep 20:597–612. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

