Advances and challenges in logical modeling of cell cycle regulation: perspective for multi-scale, integrative yeast cell models
- PMID: 27993914
- PMCID: PMC5225787
- DOI: 10.1093/femsyr/fow103
Advances and challenges in logical modeling of cell cycle regulation: perspective for multi-scale, integrative yeast cell models
Abstract
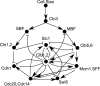
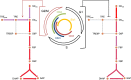
The eukaryotic cell cycle is robustly designed, with interacting molecules organized within a definite topology that ensures temporal precision of its phase transitions. Its underlying dynamics are regulated by molecular switches, for which remarkable insights have been provided by genetic and molecular biology efforts. In a number of cases, this information has been made predictive, through computational models. These models have allowed for the identification of novel molecular mechanisms, later validated experimentally. Logical modeling represents one of the youngest approaches to address cell cycle regulation. We summarize the advances that this type of modeling has achieved to reproduce and predict cell cycle dynamics. Furthermore, we present the challenge that this type of modeling is now ready to tackle: its integration with intracellular networks, and its formalisms, to understand crosstalks underlying systems level properties, ultimate aim of multi-scale models. Specifically, we discuss and illustrate how such an integration may be realized, by integrating a minimal logical model of the cell cycle with a metabolic network.
Keywords: cell cycle; constraint-based modeling; logical modeling; metabolism; multi-scale modeling; network integration.
© FEMS 2016.
Figures

Similar articles
-
Advanced Modeling of Cellular Proliferation: Toward a Multi-scale Framework Coupling Cell Cycle to Metabolism by Integrating Logical and Constraint-Based Models.Methods Mol Biol. 2019;2049:365-385. doi: 10.1007/978-1-4939-9736-7_21. Methods Mol Biol. 2019. PMID: 31602622
-
Networks and circuits in cell regulation.Biochem Biophys Res Commun. 2010 Jun 11;396(4):881-6. doi: 10.1016/j.bbrc.2010.05.015. Epub 2010 May 10. Biochem Biophys Res Commun. 2010. PMID: 20457126
-
Robustness and adaptation reveal plausible cell cycle controlling subnetwork in Saccharomyces cerevisiae.Gene. 2013 Apr 10;518(1):35-41. doi: 10.1016/j.gene.2012.11.088. Epub 2012 Dec 27. Gene. 2013. PMID: 23274654
-
From network models to network responses: integration of thermodynamic and kinetic properties of yeast genome-scale metabolic networks.FEMS Yeast Res. 2012 Mar;12(2):129-43. doi: 10.1111/j.1567-1364.2011.00771.x. Epub 2011 Dec 19. FEMS Yeast Res. 2012. PMID: 22129227 Review.
-
Cell growth and cell cycle in Saccharomyces cerevisiae: basic regulatory design and protein-protein interaction network.Biotechnol Adv. 2012 Jan-Feb;30(1):52-72. doi: 10.1016/j.biotechadv.2011.07.010. Epub 2011 Jul 22. Biotechnol Adv. 2012. PMID: 21821114 Review.
Cited by
-
Promoter-dependent nuclear RNA degradation ensures cell cycle-specific gene expression.Commun Biol. 2019 Jun 17;2:211. doi: 10.1038/s42003-019-0441-3. eCollection 2019. Commun Biol. 2019. PMID: 31240249 Free PMC article.
-
Quantitative Systems Biology to decipher design principles of a dynamic cell cycle network: the "Maximum Allowable mammalian Trade-Off-Weight" (MAmTOW).NPJ Syst Biol Appl. 2017 Sep 19;3:26. doi: 10.1038/s41540-017-0028-x. eCollection 2017. NPJ Syst Biol Appl. 2017. PMID: 28944079 Free PMC article. Review.
-
A Clb/Cdk1-mediated regulation of Fkh2 synchronizes CLB expression in the budding yeast cell cycle.NPJ Syst Biol Appl. 2017 Mar 6;3:7. doi: 10.1038/s41540-017-0008-1. eCollection 2017. NPJ Syst Biol Appl. 2017. PMID: 28649434 Free PMC article.
-
A Novel Data-Driven Boolean Model for Genetic Regulatory Networks.Front Physiol. 2018 Sep 25;9:1328. doi: 10.3389/fphys.2018.01328. eCollection 2018. Front Physiol. 2018. PMID: 30319440 Free PMC article.
-
Parameterising Translational Feedback Models of Autoregulatory RNA-Binding Proteins in Saccharomyces cerevisiae.Microorganisms. 2022 Feb 1;10(2):340. doi: 10.3390/microorganisms10020340. Microorganisms. 2022. PMID: 35208795 Free PMC article.
References
-
- Alberghina L, Mavelli G, Drovandi G et al. . Cell growth and cell cycle in Saccharomyces cerevisiae: basic regulatory design and protein–protein interaction network. Biotechnol Adv 2012;30:52–72 - PubMed
-
- Barberis M. Sic1 as a timer of Clb cyclin waves in the yeast cell cycle—design principle of not just an inhibitor. FEBS J 2012;279:3386–410 - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

