Alternative splicing and the evolution of phenotypic novelty
- PMID: 27994117
- PMCID: PMC5182408
- DOI: 10.1098/rstb.2015.0474
Alternative splicing and the evolution of phenotypic novelty
Abstract
Alternative splicing, a mechanism of post-transcriptional RNA processing whereby a single gene can encode multiple distinct transcripts, has been proposed to underlie morphological innovations in multicellular organisms. Genes with developmental functions are enriched for alternative splicing events, suggestive of a contribution of alternative splicing to developmental programmes. The role of alternative splicing as a source of transcript diversification has previously been compared to that of gene duplication, with the relationship between the two extensively explored. Alternative splicing is reduced following gene duplication with the retention of duplicate copies higher for genes which were alternatively spliced prior to duplication. Furthermore, and unlike the case for overall gene number, the proportion of alternatively spliced genes has also increased in line with the evolutionary diversification of cell types, suggesting alternative splicing may contribute to the complexity of developmental programmes. Together these observations suggest a prominent role for alternative splicing as a source of functional innovation. However, it is unknown whether the proliferation of alternative splicing events indeed reflects a functional expansion of the transcriptome or instead results from weaker selection acting on larger species, which tend to have a higher number of cell types and lower population sizes.This article is part of the themed issue 'Evo-devo in the genomics era, and the origins of morphological diversity'.
Keywords: alternative splicing; comparative genomics; effective population size; functional innovation; gene duplication; genetic drift.
© 2016 The Author(s).
Figures
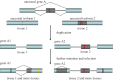
References
Publication types
MeSH terms
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources
