Characterization of a novel inhibitory human monoclonal antibody directed against Plasmodium falciparum Apical Membrane Antigen 1
- PMID: 28000709
- PMCID: PMC5175200
- DOI: 10.1038/srep39462
Characterization of a novel inhibitory human monoclonal antibody directed against Plasmodium falciparum Apical Membrane Antigen 1
Abstract
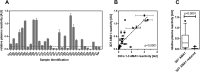
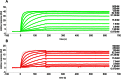
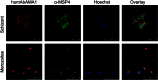
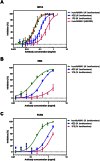
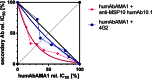
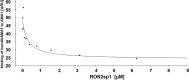
Malaria remains a major challenge to global health causing extensive morbidity and mortality. Yet, there is no efficient vaccine and the immune response remains incompletely understood. Apical Membrane Antigen 1 (AMA1), a leading vaccine candidate, plays a key role during merozoite invasion into erythrocytes by interacting with Rhoptry Neck Protein 2 (RON2). We generated a human anti-AMA1-antibody (humAbAMA1) by EBV-transformation of sorted B-lymphocytes from a Ghanaian donor and subsequent rescue of antibody variable regions. The antibody was expressed in Nicotiana benthamiana and in HEK239-6E, characterized for binding specificity and epitope, and analyzed for its inhibitory effect on Plasmodium falciparum. The generated humAbAMA1 shows an affinity of 106-135 pM. It inhibits the parasite strain 3D7A growth in vitro with an expression system-independent IC50-value of 35 μg/ml (95% confidence interval: 33 μg/ml-37 μg/ml), which is three to eight times lower than the IC50-values of inhibitory antibodies 4G2 and 1F9. The epitope was mapped to the close proximity of the RON2-peptide binding groove. Competition for binding between the RON2-peptide and humAbAMA1 was confirmed by surface plasmon resonance spectroscopy measurements. The particularly advantageous inhibitory activity of this fully human antibody might provide a basis for future therapeutic applications.
Conflict of interest statement
The authors declare no competing financial interests.
Figures


References
-
- Ramasamy R. Molecular basis for evasion of host immunity and pathogenesis in malaria. Biochim Biophys Acta 1406, 10–27 (1998). - PubMed
-
- Day K. P. & Marsh K. Naturally acquired immunity to Plasmodium falciparum. Immunology today 12, A68–71 (1991). - PubMed
-
- Langhorne J., Ndungu F. M., Sponaas A.-M. & Marsh K. Immunity to malaria: more questions than answers. Nature immunology 9, 725–732 (2008). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

