A Single Amino Acid Mutation Converts (R)-5-Diphosphomevalonate Decarboxylase into a Kinase
- PMID: 28003359
- PMCID: PMC5313113
- DOI: 10.1074/jbc.M116.752535
A Single Amino Acid Mutation Converts (R)-5-Diphosphomevalonate Decarboxylase into a Kinase
Abstract
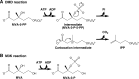
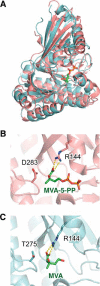
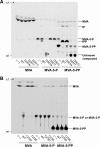
The biosynthesis of isopentenyl diphosphate, a fundamental precursor for isoprenoids, via the mevalonate pathway is completed by diphosphomevalonate decarboxylase. This enzyme catalyzes the formation of isopentenyl diphosphate through the ATP-dependent phosphorylation of the 3-hydroxyl group of (R)-5-diphosphomevalonate followed by decarboxylation coupled with the elimination of the 3-phosphate group. In this reaction, a conserved aspartate residue has been proposed to be involved in the phosphorylation step as the general base catalyst that abstracts a proton from the 3-hydroxyl group. In this study, the catalytic mechanism of this rare type of decarboxylase is re-investigated by structural and mutagenic studies on the enzyme from a thermoacidophilic archaeon Sulfolobus solfataricus The crystal structures of the archaeal enzyme in complex with (R)-5-diphosphomevalonate and adenosine 5'-O-(3-thio)triphosphate or with (R)-5-diphosphomevalonate and ADP are newly solved, and theoretical analysis based on the structure suggests the inability of proton abstraction by the conserved aspartate residue, Asp-281. Site-directed mutagenesis on Asp-281 creates mutants that only show diphosphomevalonate 3-kinase activity, demonstrating that the residue is required in the process of phosphate elimination/decarboxylation, rather than in the preceding phosphorylation step. These results enable discussion of the catalytic roles of the aspartate residue and provide clear proof of the involvement of a long predicted intermediate, (R)-3-phospho-5-diphosphomevalonate, in the reaction of the enzyme.
Keywords: archaea; crystal structure; decarboxylase; diphosphomevalonate decarboxylase; enzyme catalysis; enzyme mechanism; isoprenoid; kinase; mevalonate pathway; mutagenesis.
© 2017 by The American Society for Biochemistry and Molecular Biology, Inc.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article
Figures





References
-
- Kuzuyama T., Hemmi H., and Takahashi S. (2010) in Comprehensive Natural Products II Chemistry and Biology (Mander L., and Liu H.-W., eds) pp. 493–516, Elsevier, Oxford, United Kingdom
-
- Li T., Huo L., Pulley C., and Liu A. (2012) Decarboxylation mechanisms in biological system. Bioorg. Chem. 43, 2–14 - PubMed
-
- Froese D. S., Forouhar F., Tran T. H., Vollmar M., Kim Y. S., Lew S., Neely H., Seetharaman J., Shen Y., Xiao R., Acton T. B., Everett J. K., Cannone G., Puranik S., Savitsky P., et al. (2013) Crystal structures of malonyl-coenzyme A decarboxylase provide insights into its catalytic mechanism and disease-causing mutations. Structure 21, 1182–1192 - PMC - PubMed
-
- Bertoldi M. (2014) Mammalian dopa decarboxylase: structure, catalytic activity and inhibition. Arch. Biochem. Biophys. 546, 1–7 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

