Nuclear Localization of Integrin Cytoplasmic Domain-associated Protein-1 (ICAP1) Influences β1 Integrin Activation and Recruits Krev/Interaction Trapped-1 (KRIT1) to the Nucleus
- PMID: 28003363
- PMCID: PMC5290960
- DOI: 10.1074/jbc.M116.762393
Nuclear Localization of Integrin Cytoplasmic Domain-associated Protein-1 (ICAP1) Influences β1 Integrin Activation and Recruits Krev/Interaction Trapped-1 (KRIT1) to the Nucleus
Abstract
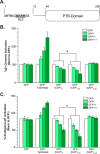
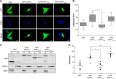
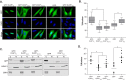
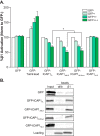
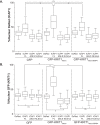
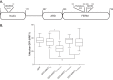
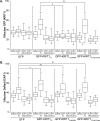
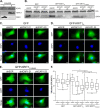
Binding of ICAP1 (integrin cytoplasmic domain-associated protein-1) to the cytoplasmic tails of β1 integrins inhibits integrin activation. ICAP1 also binds to KRIT1 (Krev interaction trapped-1), a protein whose loss of function leads to cerebral cavernous malformation, a cerebrovascular dysplasia occurring in up to 0.5% of the population. We previously showed that KRIT1 functions as a switch for β1 integrin activation by antagonizing ICAP1-mediated inhibition of integrin activation. Here we use overexpression studies, mutagenesis, and flow cytometry to show that ICAP1 contains a functional nuclear localization signal and that nuclear localization impairs the ability of ICAP1 to suppress integrin activation. Moreover, we find that ICAP1 drives the nuclear localization of KRIT1 in a manner dependent upon a direct ICAP1/KRIT1 interaction. Thus, nuclear-cytoplasmic shuttling of ICAP1 influences both integrin activation and KRIT1 localization, presumably impacting nuclear functions of KRIT1.
Keywords: ICAP1 (integrin cytoplasmic domain-associated protein); KRIT1 (Krev interaction trapped); cell compartmentalization; cerebral cavernous malformation; integrin; protein targeting.
© 2017 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures

Similar articles
-
Serine phosphorylation of the small phosphoprotein ICAP1 inhibits its nuclear accumulation.J Biol Chem. 2020 Mar 6;295(10):3269-3284. doi: 10.1074/jbc.RA119.009794. Epub 2020 Jan 31. J Biol Chem. 2020. PMID: 32005669 Free PMC article.
-
Mechanism for KRIT1 release of ICAP1-mediated suppression of integrin activation.Mol Cell. 2013 Feb 21;49(4):719-29. doi: 10.1016/j.molcel.2012.12.005. Epub 2013 Jan 11. Mol Cell. 2013. PMID: 23317506 Free PMC article.
-
Cocrystal structure of the ICAP1 PTB domain in complex with a KRIT1 peptide.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2013 May 1;69(Pt 5):494-8. doi: 10.1107/S1744309113010762. Epub 2013 Apr 27. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2013. PMID: 23695561 Free PMC article.
-
Concepts and hypothesis: integrin cytoplasmic domain-associated protein-1 (ICAP-1) as a potential player in cerebral cavernous malformation.J Neurol. 2013 Jan;260(1):10-9. doi: 10.1007/s00415-012-6567-6. Epub 2012 Jun 19. J Neurol. 2013. PMID: 22711159 Review.
-
KRIT1: A Traffic Warden at the Busy Crossroads Between Redox Signaling and the Pathogenesis of Cerebral Cavernous Malformation Disease.Antioxid Redox Signal. 2023 Mar;38(7-9):496-528. doi: 10.1089/ars.2021.0263. Epub 2022 Nov 1. Antioxid Redox Signal. 2023. PMID: 36047808 Free PMC article. Review.
Cited by
-
Chapter 22: Structural and signaling functions of integrins.Biochim Biophys Acta Biomembr. 2020 May 1;1862(5):183206. doi: 10.1016/j.bbamem.2020.183206. Epub 2020 Jan 25. Biochim Biophys Acta Biomembr. 2020. PMID: 31991120 Free PMC article. Review.
-
Working a second job: Cell adhesion proteins that moonlight in the nucleus.Front Cell Dev Biol. 2023 Apr 24;11:1163553. doi: 10.3389/fcell.2023.1163553. eCollection 2023. Front Cell Dev Biol. 2023. PMID: 37169022 Free PMC article. Review.
-
Serine phosphorylation of the small phosphoprotein ICAP1 inhibits its nuclear accumulation.J Biol Chem. 2020 Mar 6;295(10):3269-3284. doi: 10.1074/jbc.RA119.009794. Epub 2020 Jan 31. J Biol Chem. 2020. PMID: 32005669 Free PMC article.
-
KRIT1-mediated regulation of neutrophil adhesion and motility.FEBS J. 2023 Feb;290(4):1078-1095. doi: 10.1111/febs.16627. Epub 2022 Sep 20. FEBS J. 2023. PMID: 36107440 Free PMC article.
-
Is Location Everything? Regulation of the Endothelial CCM Signaling Complex.Front Cardiovasc Med. 2022 Jul 11;9:954780. doi: 10.3389/fcvm.2022.954780. eCollection 2022. Front Cardiovasc Med. 2022. PMID: 35898265 Free PMC article. Review.
References
-
- Hynes R. O. (2002) Integrins: bidirectional, allosteric signaling machines. Cell 110, 673–687 - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

