UBE3B Is a Calmodulin-regulated, Mitochondrion-associated E3 Ubiquitin Ligase
- PMID: 28003368
- PMCID: PMC5313114
- DOI: 10.1074/jbc.M116.766824
UBE3B Is a Calmodulin-regulated, Mitochondrion-associated E3 Ubiquitin Ligase
Abstract
Recent genome-wide studies found that patients with hypotonia, developmental delay, intellectual disability, congenital anomalies, characteristic facial dysmorphic features, and low cholesterol levels suffer from
Keywords: HECT; Kaufman oculocerebrofacial syndrome; calcium; mitochondria; oxidative stress; protein degradation; reactive oxygen species (ROS); super-resolution microscopy; ubiquitin/proteosome system; ubiquitylation (ubiquitination).
© 2017 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures

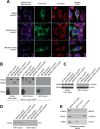
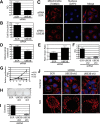

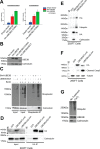

Similar articles
-
Loss of function of the E3 ubiquitin-protein ligase UBE3B causes Kaufman oculocerebrofacial syndrome.J Med Genet. 2013 Aug;50(8):493-9. doi: 10.1136/jmedgenet-2012-101405. Epub 2013 May 17. J Med Genet. 2013. PMID: 23687348 Free PMC article.
-
Expanding the clinical and mutational spectrum of Kaufman oculocerebrofacial syndrome with biallelic UBE3B mutations.Hum Genet. 2014 Jul;133(7):939-49. doi: 10.1007/s00439-014-1436-2. Epub 2014 Mar 11. Hum Genet. 2014. PMID: 24615390
-
Deficiency for the ubiquitin ligase UBE3B in a blepharophimosis-ptosis-intellectual-disability syndrome.Am J Hum Genet. 2012 Dec 7;91(6):998-1010. doi: 10.1016/j.ajhg.2012.10.011. Epub 2012 Nov 29. Am J Hum Genet. 2012. PMID: 23200864 Free PMC article.
-
Molecular Evolution, Neurodevelopmental Roles and Clinical Significance of HECT-Type UBE3 E3 Ubiquitin Ligases.Cells. 2020 Nov 10;9(11):2455. doi: 10.3390/cells9112455. Cells. 2020. PMID: 33182779 Free PMC article. Review.
-
Further phenotypic characterization of Kaufman oculocerebrofacial syndrome: report of five new cases and literature review.Clin Dysmorphol. 2019 Oct;28(4):175-183. doi: 10.1097/MCD.0000000000000282. Clin Dysmorphol. 2019. PMID: 31162149 Review.
Cited by
-
NAD+ bioavailability mediates PARG inhibition-induced replication arrest, intra S-phase checkpoint and apoptosis in glioma stem cells.NAR Cancer. 2021 Nov 17;3(4):zcab044. doi: 10.1093/narcan/zcab044. eCollection 2021 Dec. NAR Cancer. 2021. PMID: 34806016 Free PMC article.
-
Genome-Wide Association Analysis of Boar Semen Traits Based on Computer-Assisted Semen Analysis and Flow Cytometry.Animals (Basel). 2024 Dec 26;15(1):26. doi: 10.3390/ani15010026. Animals (Basel). 2024. PMID: 39794969 Free PMC article.
-
The N-cadherin interactome in primary cardiomyocytes as defined using quantitative proximity proteomics.J Cell Sci. 2019 Feb 11;132(3):jcs221606. doi: 10.1242/jcs.221606. J Cell Sci. 2019. PMID: 30630894 Free PMC article.
-
Nrf2/UBE3B protects against acute lung injury by inhibiting ferritinophagy through the ubiquitination of NCOA4.Biol Direct. 2025 Jul 16;20(1):85. doi: 10.1186/s13062-025-00678-z. Biol Direct. 2025. PMID: 40671136 Free PMC article.
-
oxi-1 and fshr-1 are required for neuromuscular signaling under normal and oxidative stress conditions in C. elegans.MicroPubl Biol. 2018 Aug 28;2018:10.17912/pfyw-ft85. doi: 10.17912/pfyw-ft85. MicroPubl Biol. 2018. PMID: 32550383 Free PMC article. No abstract available.
References
-
- Komander D. (2009) The emerging complexity of protein ubiquitination. Biochem. Soc. Trans. 37, 937–953 - PubMed
-
- Tai H. C., and Schuman E. M. (2008) Ubiquitin, the proteasome and protein degradation in neuronal function and dysfunction. Nat. Rev. Neurosci. 9, 826–838 - PubMed
-
- Kawabe H., and Brose N. (2011) The role of ubiquitylation in nerve cell development. Nat. Rev. Neurosci. 12, 251–268 - PubMed
-
- Varshavsky A. (2012) The ubiquitin system, an immense realm. Annu. Rev. Biochem. 81, 167–176 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

