Spatially-Resolved Proteomics: Rapid Quantitative Analysis of Laser Capture Microdissected Alveolar Tissue Samples
- PMID: 28004771
- PMCID: PMC5177886
- DOI: 10.1038/srep39223
Spatially-Resolved Proteomics: Rapid Quantitative Analysis of Laser Capture Microdissected Alveolar Tissue Samples
Abstract
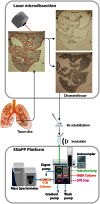
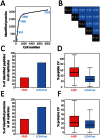
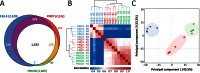
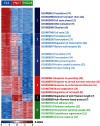
Laser capture microdissection (LCM)-enabled region-specific tissue analyses are critical to better understand complex multicellular processes. However, current proteomics workflows entail several manual sample preparation steps and are challenged by the microscopic mass-limited samples generated by LCM, impacting measurement robustness, quantification and throughput. Here, we coupled LCM with a proteomics workflow that provides fully automated analysis of proteomes from microdissected tissues. Benchmarking against the current state-of-the-art in ultrasensitive global proteomics (FASP workflow), our approach demonstrated significant improvements in quantification (~2-fold lower variance) and throughput (>5 times faster). Using our approach we for the first time characterized, to a depth of >3,400 proteins, the ontogeny of protein changes during normal lung development in microdissected alveolar tissue containing only 4,000 cells. Our analysis revealed seven defined modules of coordinated transcription factor-signaling molecule expression patterns, suggesting a complex network of temporal regulatory control directs normal lung development with epigenetic regulation fine-tuning pre-natal developmental processes.
Conflict of interest statement
The authors declare no competing financial interests.
Figures


References
-
- Larance M. & Lamond A. I. Multidimensional proteomics for cell biology. Nat. Rev. Mol. Cell Biol. 16, 269–280 (2015). - PubMed
-
- Longuespée R. et al.. Tissue proteomics for the next decade? Towards a molecular dimension in histology. OMICS 18, 539–52 (2014). - PubMed
-
- Heeren R. M. A., Smith D. F., Stauber J., Kükrer-Kaletas B. & MacAleese L. Imaging mass spectrometry: hype or hope? J. Am. Soc. Mass Spectrom. 20, 1006–14 (2009). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

