Coupled ATPase-adenylate kinase activity in ABC transporters
- PMID: 28004795
- PMCID: PMC5192220
- DOI: 10.1038/ncomms13864
Coupled ATPase-adenylate kinase activity in ABC transporters
Abstract
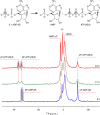
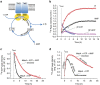
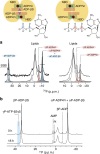
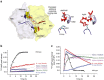
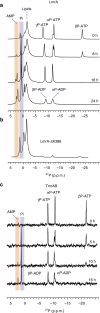
ATP-binding cassette (ABC) transporters, a superfamily of integral membrane proteins, catalyse the translocation of substrates across the cellular membrane by ATP hydrolysis. Here we demonstrate by nucleotide turnover and binding studies based on 31P solid-state NMR spectroscopy that the ABC exporter and lipid A flippase MsbA can couple ATP hydrolysis to an adenylate kinase activity, where ADP is converted into AMP and ATP. Single-point mutations reveal that both ATPase and adenylate kinase mechanisms are associated with the same conserved motifs of the nucleotide-binding domain. Based on these results, we propose a model for the coupled ATPase-adenylate kinase mechanism, involving the canonical and an additional nucleotide-binding site. We extend these findings to other prokaryotic ABC exporters, namely LmrA and TmrAB, suggesting that the coupled activities are a general feature of ABC exporters.
Figures


Similar articles
-
Probing native metal ion association sites through quenching of fluorophores in the nucleotide-binding domains of the ABC transporter MsbA.Biochem J. 2017 May 30;474(12):1993-2007. doi: 10.1042/BCJ20161051. Biochem J. 2017. PMID: 28432259
-
Structural basis for adenylate kinase activity in ABC ATPases.J Mol Biol. 2010 Aug 13;401(2):265-73. doi: 10.1016/j.jmb.2010.06.029. Epub 2010 Jun 19. J Mol Biol. 2010. PMID: 20600125
-
Functional characterization and ATP-induced dimerization of the isolated ABC-domain of the haemolysin B transporter.Biochemistry. 2005 Jul 19;44(28):9680-90. doi: 10.1021/bi0506122. Biochemistry. 2005. PMID: 16008353
-
Structural basis for the mechanism of ABC transporters.Biochem Soc Trans. 2015 Oct;43(5):889-93. doi: 10.1042/BST20150047. Biochem Soc Trans. 2015. PMID: 26517899 Review.
-
The role of the degenerate nucleotide binding site in type I ABC exporters.FEBS Lett. 2020 Dec;594(23):3815-3838. doi: 10.1002/1873-3468.13997. Epub 2020 Nov 27. FEBS Lett. 2020. PMID: 33179257 Free PMC article. Review.
Cited by
-
Detection of side-chain proton resonances of fully protonated biosolids in nano-litre volumes by magic angle spinning solid-state NMR.J Biomol NMR. 2018 Mar;70(3):177-185. doi: 10.1007/s10858-018-0168-3. Epub 2018 Mar 3. J Biomol NMR. 2018. PMID: 29502224
-
A New Method of Assessing Lipid Mixtures by 31P Magic-Angle Spinning NMR.Biophys J. 2018 Mar 27;114(6):1368-1376. doi: 10.1016/j.bpj.2018.01.025. Biophys J. 2018. PMID: 29590594 Free PMC article.
-
Easy Synthesis of Complex Biomolecular Assemblies: Wheat Germ Cell-Free Protein Expression in Structural Biology.Front Mol Biosci. 2021 Mar 25;8:639587. doi: 10.3389/fmolb.2021.639587. eCollection 2021. Front Mol Biosci. 2021. PMID: 33842544 Free PMC article. Review.
-
Structural basis for lipid and copper regulation of the ABC transporter MsbA.Nat Commun. 2022 Nov 26;13(1):7291. doi: 10.1038/s41467-022-34905-2. Nat Commun. 2022. PMID: 36435815 Free PMC article.
-
Interaction of a Homologous Series of Amphiphiles with P-glycoprotein in a Membrane Environment-Contributions of Polar and Non-Polar Interactions.Pharmaceutics. 2023 Jan 3;15(1):174. doi: 10.3390/pharmaceutics15010174. Pharmaceutics. 2023. PMID: 36678803 Free PMC article.
References
-
- Higgins C. F. ABC transporters: from microorganisms to man. Annu. Rev. Cell Biol. 8, 67–113 (1992). - PubMed
-
- Higgins C. F. Multiple molecular mechanisms for multidrug resistance transporters. Nature 446, 749–757 (2007). - PubMed
-
- Dawson R. J. & Locher K. P. Structure of a bacterial multidrug ABC transporter. Nature 443, 180–185 (2006). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

