Probing Enhanced Double-Strand Break Formation at Abasic Sites within Clustered Lesions in Nucleosome Core Particles
- PMID: 28005342
- PMCID: PMC5372979
- DOI: 10.1021/acs.biochem.6b01144
Probing Enhanced Double-Strand Break Formation at Abasic Sites within Clustered Lesions in Nucleosome Core Particles
Abstract
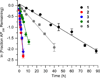
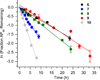
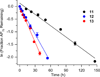
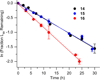
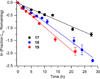
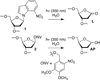
DNA is rapidly cleaved under mild alkaline conditions at apyrimidinic/apurinic sites, but the half-life is several weeks in phosphate buffer (pH 7.5). However, abasic sites are ∼100-fold more reactive within nucleosome core particles (NCPs). Histone proteins catalyze the strand scission, and at superhelical location 1.5, the histone H4 tail is largely responsible for the accelerated cleavage. The rate constant for strand scission at an abasic site is enhanced further in a nucleosome core particle when it is part of a bistranded lesion containing a proximal strand break. Cleavage of this form results in a highly deleterious double-strand break. This acceleration is dependent upon the position of the abasic lesion in the NCP and its structure. The enhancement in cleavage rate at an apurinic/apyrimidinic site rapidly drops off as the distance between the strand break and abasic site increases and is negligible once the two forms of damage are separated by 7 bp. However, the enhancement of the rate of double-strand break formation increases when the size of the gap is increased from one to two nucleotides. In contrast, the cleavage rate enhancement at 2-deoxyribonolactone within bistranded lesions is more modest, and it is similar in free DNA and nucleosome core particles. We postulate that the enhanced rate of double-strand break formation at bistranded lesions containing apurinic/apyrimidinic sites within nucleosome core particles is a general phenomenon and is due to increased DNA flexibility.
Figures

References
-
- Chapman JR, Taylor MRG, Boulton SJ. Playing the end game: DNA double-strand break repair pathway choice. Mol. Cell. 2012;47:497–510. - PubMed
-
- Srivastava M, Raghavan SC. DNA double-strand break repair inhibitors as cancer therapeutics. Chem. Biol. 2015;22:17–29. - PubMed
-
- Srivastava M, Nambiar M, Sharma S, Karki SS, Goldsmith G, Hegde M, Kumar S, Pandey M, Singh RK, Ray P, Natarajan R, Kelkar M, De A, Choudhary B, Raghavan SC. An inhibitor of nonhomologous end-joining abrogates double-strand break repair and impedes cancer progression. Cell. 2012;151:1474–1487. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

