Implementation of next generation sequencing into pediatric hematology-oncology practice: moving beyond actionable alterations
- PMID: 28007021
- PMCID: PMC5180407
- DOI: 10.1186/s13073-016-0389-6
Implementation of next generation sequencing into pediatric hematology-oncology practice: moving beyond actionable alterations
Abstract
Background: Molecular characterization has the potential to advance the management of pediatric cancer and high-risk hematologic disease. The clinical integration of genome sequencing into standard clinical practice has been limited and the potential utility of genome sequencing to identify clinically impactful information beyond targetable alterations has been underestimated.
Methods: The Precision in Pediatric Sequencing (PIPseq) Program at Columbia University Medical Center instituted prospective clinical next generation sequencing (NGS) for pediatric cancer and hematologic disorders at risk for treatment failure. We performed cancer whole exome sequencing (WES) of patient-matched tumor-normal samples and RNA sequencing (RNA-seq) of tumor to identify sequence variants, fusion transcripts, relative gene expression, and copy number variation (CNV). A directed cancer gene panel assay was used when sample adequacy was a concern. Constitutional WES of patients and parents was performed when a constitutionally encoded disease was suspected. Results were initially reviewed by a molecular pathologist and subsequently by a multi-disciplinary molecular tumor board. Clinical reports were issued to the ordering physician and posted to the patient's electronic medical record.
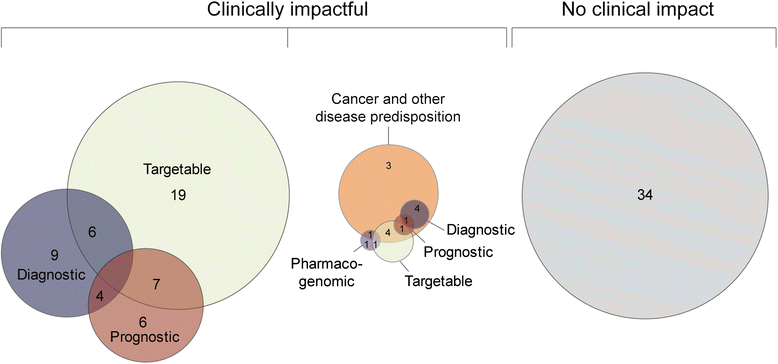
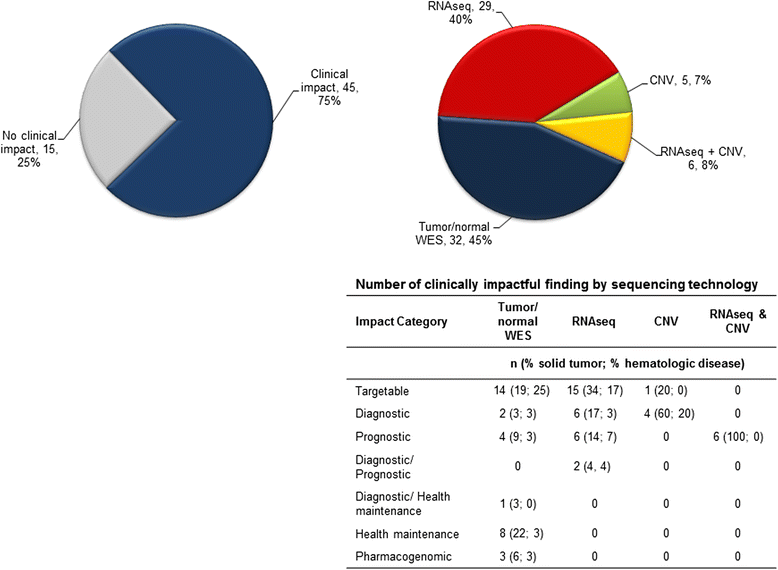
Results: NGS was performed on tumor and/or normal tissue from 101 high-risk pediatric patients. Potentially actionable alterations were identified in 38% of patients, of which only 16% subsequently received matched therapy. In an additional 38% of patients, the genomic data provided clinically relevant information of diagnostic, prognostic, or pharmacogenomic significance. RNA-seq was clinically impactful in 37/65 patients (57%) providing diagnostic and/or prognostic information for 17 patients (26%) and identified therapeutic targets in 15 patients (23%). Known or likely pathogenic germline alterations were discovered in 18/90 patients (20%) with 14% having germline alternations in cancer predisposition genes. American College of Medical Genetics (ACMG) secondary findings were identified in six patients.
Conclusions: Our results demonstrate the feasibility of incorporating clinical NGS into pediatric hematology-oncology practice. Beyond the identification of actionable alterations, the ability to avoid ineffective/inappropriate therapies, make a definitive diagnosis, and identify pharmacogenomic modifiers is clinically impactful. Taking a more inclusive view of potential clinical utility, 66% of cases tested through our program had clinically impactful findings and samples interrogated with both WES and RNA-seq resulted in data that impacted clinical decisions in 75% of cases.
Keywords: Pediatric oncology; Precision medicine; RNA sequencing; Whole exome sequencing.
Figures



References
-
- Murphy S, Xu J, Kochanek K. Deaths: Final Data for 2010. National vital statistics reports, vol. 61 no 4. Hyattsville, MD: National Center for Health Statistics; 2013. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

