Critical role of the peroxisomal protein PEX16 in white adipocyte development and lipid homeostasis
- PMID: 28017862
- PMCID: PMC7116240
- DOI: 10.1016/j.bbalip.2016.12.009
Critical role of the peroxisomal protein PEX16 in white adipocyte development and lipid homeostasis
Abstract
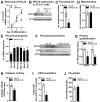
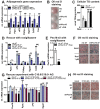
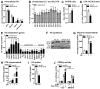
The importance of peroxisomes for adipocyte function is poorly understood. Herein, we provide insights into the critical role of peroxin 16 (PEX16)-mediated peroxisome biogenesis in adipocyte development and lipid metabolism. Pex16 is highly expressed in adipose tissues and upregulated during adipogenesis of murine and human cells. We demonstrate that Pex16 is a target gene of the adipogenesis "master-regulator" PPARγ. Stable silencing of Pex16 in 3T3-L1 cells strongly reduced the number of peroxisomes while mitochondrial number was unaffected. Concomitantly, peroxisomal fatty acid (FA) oxidation was reduced, thereby causing accumulation of long- and very long-chain (polyunsaturated) FAs and reduction of odd-chain FAs. Further, Pex16-silencing decreased cellular oxygen consumption and increased FA release. Additionally, silencing of Pex16 impaired adipocyte differentiation, lipogenic and adipogenic marker gene expression, and cellular triglyceride stores. Addition of PPARγ agonist rosiglitazone and peroxisome-related lipid species to Pex16-silenced 3T3-L1 cells rescued adipogenesis. These data provide evidence that PEX16 is required for peroxisome biogenesis and highlights the relevance of peroxisomes for adipogenesis and adipocyte lipid metabolism.
Keywords: Adipogenesis; Lipid homeostasis; PEX16; PPARγ; Peroxisome.
Copyright © 2016 The Author(s). Published by Elsevier B.V. All rights reserved.
Conflict of interest statement
The authors declare no potential conflict of interest.
Figures


Similar articles
-
Coordinate functional regulation between microsomal prostaglandin E synthase-1 (mPGES-1) and peroxisome proliferator-activated receptor γ (PPARγ) in the conversion of white-to-brown adipocytes.J Biol Chem. 2013 Sep 27;288(39):28230-42. doi: 10.1074/jbc.M113.468603. Epub 2013 Aug 13. J Biol Chem. 2013. PMID: 23943621 Free PMC article.
-
Hepatocyte nuclear factor 1b is a novel negative regulator of white adipocyte differentiation.Cell Death Differ. 2017 Sep;24(9):1588-1597. doi: 10.1038/cdd.2017.85. Epub 2017 Jun 16. Cell Death Differ. 2017. PMID: 28622294 Free PMC article.
-
Obestatin and Rosiglitazone Differentially Modulate Lipid Metabolism Through Peroxisome Proliferator-activated Receptor-γ (PPARγ) in Pre-adipose and Mature 3T3-L1 Cells.Cell Biochem Biophys. 2021 Mar;79(1):73-85. doi: 10.1007/s12013-020-00958-7. Epub 2021 Jan 11. Cell Biochem Biophys. 2021. PMID: 33432549
-
Peroxisomal regulation of redox homeostasis and adipocyte metabolism.Redox Biol. 2019 Jun;24:101167. doi: 10.1016/j.redox.2019.101167. Epub 2019 Mar 14. Redox Biol. 2019. PMID: 30921635 Free PMC article. Review.
-
Molecular mechanism of down-regulating adipogenic transcription factors in 3T3-L1 adipocyte cells by bioactive anti-adipogenic compounds.Mol Biol Rep. 2021 Jan;48(1):743-761. doi: 10.1007/s11033-020-06036-8. Epub 2020 Dec 4. Mol Biol Rep. 2021. PMID: 33275195 Review.
Cited by
-
1-O-Alkylglycerol Ethers from the Marine Sponge Guitarra abbotti and Their Cytotoxic Activity.Mar Drugs. 2022 Jun 22;20(7):409. doi: 10.3390/md20070409. Mar Drugs. 2022. PMID: 35877702 Free PMC article.
-
Adaptations of the 3T3-L1 adipocyte lipidome to defective ether lipid catabolism upon Agmo knockdown.J Lipid Res. 2022 Jun;63(6):100222. doi: 10.1016/j.jlr.2022.100222. Epub 2022 May 7. J Lipid Res. 2022. PMID: 35537527 Free PMC article.
-
Analysis of the differentially expressed genes in the combs and testes of Qingyuan partridge roosters at different developmental stages.BMC Genomics. 2024 Jan 4;25(1):33. doi: 10.1186/s12864-024-09960-2. BMC Genomics. 2024. PMID: 38177997 Free PMC article.
-
Weighted Single-Step GWAS Identified Candidate Genes Associated with Growth Traits in a Duroc Pig Population.Genes (Basel). 2021 Jan 19;12(1):117. doi: 10.3390/genes12010117. Genes (Basel). 2021. PMID: 33477978 Free PMC article.
-
Peroxisomal Stress Response and Inter-Organelle Communication in Cellular Homeostasis and Aging.Antioxidants (Basel). 2022 Jan 19;11(2):192. doi: 10.3390/antiox11020192. Antioxidants (Basel). 2022. PMID: 35204075 Free PMC article. Review.
References
-
- Hakim-Weber R, Krogsdam A-M, Jørgensen C, Fischer M, Prokesch A, Bogner-Strauss JG, et al. Transcriptional regulatory program in wild-type and retinoblastoma gene-deficient mouse embryonic fibroblasts during adipocyte differentiation. BMC Res Notes. 2011;4(1):157. doi: 10.1186/1756-0500-4-157. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

