CRISPR-Mediated Epigenome Editing
- PMID: 28018139
- PMCID: PMC5168826
CRISPR-Mediated Epigenome Editing
Abstract
Mounting evidence has called into question our understanding of the role that the central dogma of molecular biology plays in human pathology. The conventional view that elucidating the mechanisms for translating genes into proteins can account for a panoply of diseases has proven incomplete. Landmark studies point to epigenetics as a missing piece of the puzzle. However, technological limitations have hindered the study of specific roles for histone post-translational modifications, DNA modifications, and non-coding RNAs in regulation of the epigenome and chromatin structure. This feature highlights CRISPR systems, including CRISPR-Cas9, as novel tools for targeted epigenome editing. It summarizes recent developments in the field, including integration of optogenetic and functional genomic approaches to explore new therapeutic opportunities, and underscores the importance of mitigating current limitations in the field. This comprehensive, analytical assessment identifies current research gaps, forecasts future research opportunities, and argues that as epigenome editing technologies mature, overcoming critical challenges in delivery, specificity, and fidelity should clear the path to bring these technologies into the clinic.
Keywords: CRISPR; CRISPR systems for epigenome editing; CRISPR-Cas9; Epigenome editing; epigenetics; epigenome engineering; histone and DNA epigenetic modifications; optogenetics; transcription activation; transcription repression.
Figures
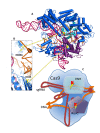

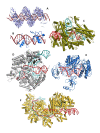
Similar articles
-
Epigenome editing by CRISPR/Cas9 in clinical settings: possibilities and challenges.Brief Funct Genomics. 2020 May 20;19(3):215-228. doi: 10.1093/bfgp/elz035. Brief Funct Genomics. 2020. PMID: 31819946 Review.
-
Zinc Fingers, TALEs, and CRISPR Systems: A Comparison of Tools for Epigenome Editing.Methods Mol Biol. 2018;1767:19-63. doi: 10.1007/978-1-4939-7774-1_2. Methods Mol Biol. 2018. PMID: 29524128 Review.
-
Investigating crosstalk between H3K27 acetylation and H3K4 trimethylation in CRISPR/dCas-based epigenome editing and gene activation.Sci Rep. 2021 Aug 5;11(1):15912. doi: 10.1038/s41598-021-95398-5. Sci Rep. 2021. PMID: 34354157 Free PMC article.
-
CRISPR/Cas9 guided genome and epigenome engineering and its therapeutic applications in immune mediated diseases.Semin Cell Dev Biol. 2019 Dec;96:32-43. doi: 10.1016/j.semcdb.2019.05.007. Epub 2019 Jun 20. Semin Cell Dev Biol. 2019. PMID: 31112800 Review.
-
Epigenomics-Guided Drug Development: Recent Advances in Solving the Cancer Treatment "jigsaw puzzle".OMICS. 2019 Feb;23(2):70-85. doi: 10.1089/omi.2018.0206. OMICS. 2019. PMID: 30767728
Cited by
-
The Relevance of Gender in Tumor-Influencing Epigenetic Traits.Epigenomes. 2019 Jan 28;3(1):6. doi: 10.3390/epigenomes3010006. Epigenomes. 2019. PMID: 34991275 Free PMC article. Review.
-
Current Bioinformatics Tools to Optimize CRISPR/Cas9 Experiments to Reduce Off-Target Effects.Int J Mol Sci. 2023 Mar 27;24(7):6261. doi: 10.3390/ijms24076261. Int J Mol Sci. 2023. PMID: 37047235 Free PMC article.
-
Divergent susceptibilities to AAV-SaCas9-gRNA vector-mediated genome-editing in a single-cell-derived cell population.BMC Res Notes. 2017 Dec 8;10(1):720. doi: 10.1186/s13104-017-3028-4. BMC Res Notes. 2017. PMID: 29221488 Free PMC article.
-
Epigenetic inactivation of tumour suppressor coding and non-coding genes in human cancer: an update.Open Biol. 2017 Sep;7(9):170152. doi: 10.1098/rsob.170152. Open Biol. 2017. PMID: 28931650 Free PMC article. Review.
-
Elevated BICD2 DNA methylation in blood of major depressive disorder patients and reduction of depressive-like behaviors in hippocampal Bicd2-knockdown mice.Proc Natl Acad Sci U S A. 2022 Jul 26;119(30):e2201967119. doi: 10.1073/pnas.2201967119. Epub 2022 Jul 18. Proc Natl Acad Sci U S A. 2022. PMID: 35858435 Free PMC article.
References
-
- Waddington CH. Genetic Assimilation of the Bithorax Phenotype. Evolution. 1956;10(1):1–13.
-
- Waddington CH. The Strategy of the Genes. London: George Allen & Unwin; 1957.
-
- Goldberg AD. et al. Epigenetics: a landscape takes shape. Cell. 2007;128(4):635–638. - PubMed
-
- Young IT. et al. Characterization of chromatin distribution in cell nuclei. Cytometry. 1986;7(5):467–474. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
