Genome-wide mapping of autonomous promoter activity in human cells
- PMID: 28024146
- PMCID: PMC5498152
- DOI: 10.1038/nbt.3754
Genome-wide mapping of autonomous promoter activity in human cells
Abstract
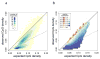
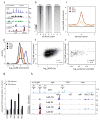
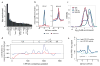
Previous methods to systematically characterize sequence-intrinsic activity of promoters have been limited by relatively low throughput and the length of the sequences that could be tested. Here we present 'survey of regulatory elements' (SuRE), a method that assays more than 108 DNA fragments, each 0.2-2 kb in size, for their ability to drive transcription autonomously. In SuRE, a plasmid library of random genomic fragments upstream of a 20-bp barcode is constructed, and decoded by paired-end sequencing. This library is used to transfect cells, and barcodes in transcribed RNA are quantified by high-throughput sequencing. When applied to the human genome, we achieve 55-fold genome coverage, allowing us to map autonomous promoter activity genome-wide in K562 cells. By computational modeling we delineate subregions within promoters that are relevant for their activity. We show that antisense promoter transcription is generally dependent on the sense core promoter sequences, and that most enhancers and several families of repetitive elements act as autonomous transcription initiation sites.
Conflict of interest statement
The authors declare no competing financial interest.
Figures

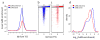

Comment in
-
Core promoters across the genome.Nat Biotechnol. 2017 Feb 8;35(2):123-124. doi: 10.1038/nbt.3788. Nat Biotechnol. 2017. PMID: 28178253 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

