Specific regions of the brain are capable of fructose metabolism
- PMID: 28034722
- PMCID: PMC5420427
- DOI: 10.1016/j.brainres.2016.12.022
Specific regions of the brain are capable of fructose metabolism
Abstract
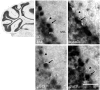
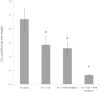
High fructose consumption in the Western diet correlates with disease states such as obesity and metabolic syndrome complications, including type II diabetes, chronic kidney disease, and non-alcoholic fatty acid liver disease. Liver and kidneys are responsible for metabolism of 40-60% of ingested fructose, while the physiological fate of the remaining fructose remains poorly understood. The primary metabolic pathway for fructose includes the fructose-transporting solute-like carrier transport proteins 2a (SLC2a or GLUT), including GLUT5 and GLUT9, ketohexokinase (KHK), and aldolase. Bioinformatic analysis of gene expression encoding these proteins (glut5, glut9, khk, and aldoC, respectively) identifies other organs capable of this fructose metabolism. This analysis predicts brain, lymphoreticular tissue, placenta, and reproductive tissues as possible additional organs for fructose metabolism. While expression of these genes is highest in liver, the brain is predicted to have expression levels of these genes similar to kidney. RNA in situ hybridization of coronal slices of adult mouse brains validate the in silico expression of glut5, glut9, khk, and aldoC, and show expression across many regions of the brain, with the most notable expression in the cerebellum, hippocampus, cortex, and olfactory bulb. Dissected samples of these brain regions show KHK and aldolase enzyme activity 5-10 times the concentration of that in liver. Furthermore, rates of fructose oxidation in these brain regions are 15-150 times that of liver slices, confirming the bioinformatics prediction and in situ hybridization data. This suggests that previously unappreciated regions across the brain can use fructose, in addition to glucose, for energy production.
Keywords: Brain energy metabolism; Fructose toxicity; Fructose transporters.
Copyright © 2016 Elsevier B.V. All rights reserved.
Figures




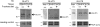
References
-
- Abbott LC, Isaacs KR, Heckroth JA. Co-localization of tyrosine hydroxylase and zebrin II immunoreactivities in Purkinje cells of the mutant mice, tottering and tottering/leaner. Neurosci. 1996;71:461–75. - PubMed
-
- Adelman RC, Ballard FJ, Weinhouse S. Purification and properties of rat liver fructokinase. J Biol Chem. 1967;242:3360–5. - PubMed
-
- Ahn AH, Dziennis S, Hawkes R, Herrup K. The cloning of zebrin II reveals its identity with aldolase C. Development. 1994;120:2081–90. - PubMed
-
- Anzai N, Ichida K, Jutabha P, Kimura T, Babu E, Jin CJ, Srivastava S, Kitamura K, Hisatome I, Endou H, Sakurai H. Plasma urate level is directly regulated by a voltage-driven urate efflux transporter URATV1 (SLC2a9) in humans. J Biol Chem. 2008;283:26834–8. - PubMed
-
- Augustin R, Carayannopoulos MO, Dowd LO, Phay JE, Moley JF, Moley KH. Identification and characterization of human glucose transporter-like protein-9 (GLUT9): Alternative splicing alters trafficking. J Biol Chem. 2004;279:16229–36. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

