The metastasis suppressor RARRES3 as an endogenous inhibitor of the immunoproteasome expression in breast cancer cells
- PMID: 28051153
- PMCID: PMC5209724
- DOI: 10.1038/srep39873
The metastasis suppressor RARRES3 as an endogenous inhibitor of the immunoproteasome expression in breast cancer cells
Abstract
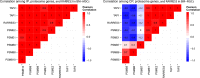
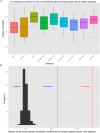
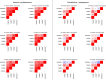
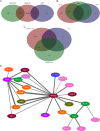
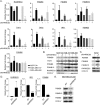
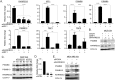
In breast cancer metastasis, the dynamic continuum involving pro- and anti-inflammatory regulators can become compromised. Over 600 genes have been implicated in metastasis to bone, lung or brain but how these genes might contribute to perturbation of immune function is poorly understood. To gain insight, we adopted a gene co-expression network approach that draws on the functional parallels between naturally occurring bone marrow-derived mesenchymal stem cells (BM-MSCs) and cancer stem cells (CSCs). Our network analyses indicate a key role for metastasis suppressor RARRES3, including potential to regulate the immunoproteasome (IP), a specialized proteasome induced under inflammatory conditions. Knockdown of RARRES3 in near-normal mammary epithelial and breast cancer cell lines increases overall transcript and protein levels of the IP subunits, but not of their constitutively expressed counterparts. RARRES3 mRNA expression is controlled by interferon regulatory factor IRF1, an inducer of the IP, and is sensitive to depletion of the retinoid-related receptor RORA that regulates various physiological processes including immunity through modulation of gene expression. Collectively, these findings identify a novel regulatory role for RARRES3 as an endogenous inhibitor of IP expression, and contribute to our evolving understanding of potential pathways underlying breast cancer driven immune modulation.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

