Histone chaperone networks shaping chromatin function
- PMID: 28053344
- PMCID: PMC5319910
- DOI: 10.1038/nrm.2016.159
Histone chaperone networks shaping chromatin function
Abstract
The association of histones with specific chaperone complexes is important for their folding, oligomerization, post-translational modification, nuclear import, stability, assembly and genomic localization. In this way, the chaperoning of soluble histones is a key determinant of histone availability and fate, which affects all chromosomal processes, including gene expression, chromosome segregation and genome replication and repair. Here, we review the distinct structural and functional properties of the expanding network of histone chaperones. We emphasize how chaperones cooperate in the histone chaperone network and via co-chaperone complexes to match histone supply with demand, thereby promoting proper nucleosome assembly and maintaining epigenetic information by recycling modified histones evicted from chromatin.
Figures



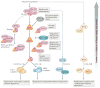

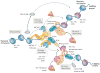
References
-
- Laskey R, Honda B, Mills A, Finch J. Nucleosomes are assembled by an acidic protein which binds histones and transfers them to DNA. Nature. 1978;275:416–420. The first description of histone chaperone function elucidated by classical biochemical approaches probing the physicochemical nature of the association between nucleoplasmin and histones. - PubMed
-
- Venkatesh S, Workman JL. Histone exchange, chromatin structure and the regulation of transcription. Nat Rev Mol Cell Biol. 2015;16:178–189. - PubMed
-
- Alabert C, Groth A. Chromatin replication and epigenome maintenance. Nat Rev Mol Cell Biol. 2012;13:153–167. - PubMed
-
- Annunziato AT. Assembling chromatin: the long and winding road. Biochim Biophys Acta. 2013;1819:196–210. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

