Coordinated regulation of acid resistance in Escherichia coli
- PMID: 28061857
- PMCID: PMC5217608
- DOI: 10.1186/s12918-016-0376-y
Coordinated regulation of acid resistance in Escherichia coli
Abstract
Background: Enteric Escherichia coli survives the highly acidic environment of the stomach through multiple acid resistance (AR) mechanisms. The most effective system, AR2, decarboxylates externally-derived glutamate to remove cytoplasmic protons and excrete GABA. The first described system, AR1, does not require an external amino acid. Its mechanism has not been determined. The regulation of the multiple AR systems and their coordination with broader cellular metabolism has not been fully explored.
Results: We utilized a combination of ChIP-Seq and gene expression analysis to experimentally map the regulatory interactions of four TFs: nac, ntrC, ompR, and csiR. Our data identified all previously in vivo confirmed direct interactions and revealed several others previously inferred from gene expression data. Our data demonstrate that nac and csiR directly modulate AR, and leads to a regulatory network model in which all four TFs participate in coordinating acid resistance, glutamate metabolism, and nitrogen metabolism. This model predicts a novel mechanism for AR1 by which the decarboxylation enzymes of AR2 are used with internally derived glutamate. This hypothesis makes several testable predictions that we confirmed experimentally.
Conclusions: Our data suggest that the regulatory network underlying AR is complex and deeply interconnected with the regulation of GABA and glutamate metabolism, nitrogen metabolism. These connections underlie and experimentally validated model of AR1 in which the decarboxylation enzymes of AR2 are used with internally derived glutamate.
Keywords: Acid resistance; Regulatory network modeling; Systems biology/ChIP-Seq.
Figures
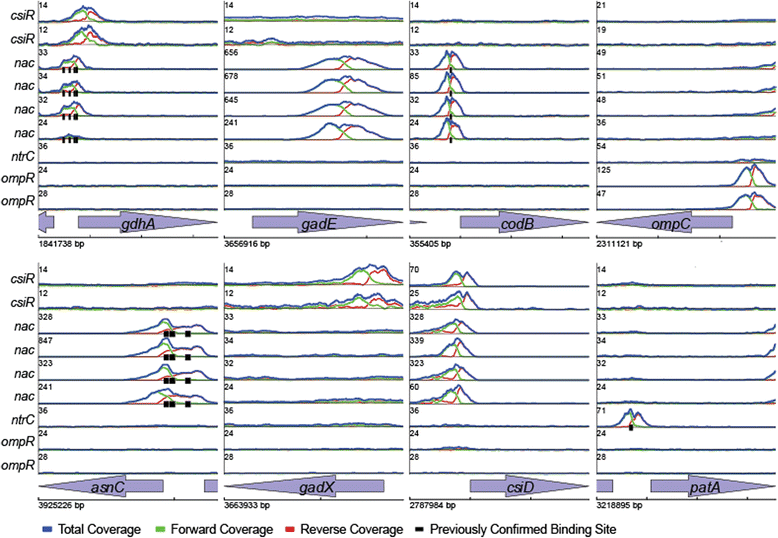


References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

