Redox crisis underlies conditional light-dark lethality in cyanobacterial mutants that lack the circadian regulator, RpaA
- PMID: 28074036
- PMCID: PMC5278464
- DOI: 10.1073/pnas.1613078114
Redox crisis underlies conditional light-dark lethality in cyanobacterial mutants that lack the circadian regulator, RpaA
Abstract
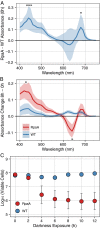
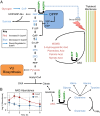
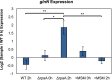
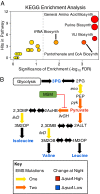
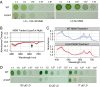
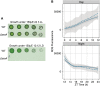
Cyanobacteria evolved a robust circadian clock, which has a profound influence on fitness and metabolism under daily light-dark (LD) cycles. In the model cyanobacterium Synechococcus elongatus PCC 7942, a functional clock is not required for diurnal growth, but mutants defective for the response regulator that mediates transcriptional rhythms in the wild-type, regulator of phycobilisome association A (RpaA), cannot be cultured under LD conditions. We found that rpaA-null mutants are inviable after several hours in the dark and compared the metabolomes of wild-type and rpaA-null strains to identify the source of lethality. Here, we show that the wild-type metabolome is very stable throughout the night, and this stability is lost in the absence of RpaA. Additionally, an rpaA mutant accumulates excessive reactive oxygen species (ROS) during the day and is unable to clear it during the night. The rpaA-null metabolome indicates that these cells are reductant-starved in the dark, likely because enzymes of the primary nighttime NADPH-producing pathway are direct targets of RpaA. Because NADPH is required for processes that detoxify ROS, conditional LD lethality likely results from inability of the mutant to activate reductant-requiring pathways that detoxify ROS when photosynthesis is not active. We identified second-site mutations and growth conditions that suppress LD lethality in the mutant background that support these conclusions. These results provide a mechanistic explanation as to why rpaA-null mutants die in the dark, further connect the clock to metabolism under diurnal growth, and indicate that RpaA likely has important unidentified functions during the day.
Keywords: circadian clock; cyanobacteria; diurnal; metabolism; metabolomics.
Conflict of interest statement
The authors declare no conflict of interest. Dedicated to the memory of Dr. David B. Knaff.
Figures








References
-
- Oliver JW, Atsumi S. Metabolic design for cyanobacterial chemical synthesis. Photosynth Res. 2014;120(3):249–261. - PubMed
-
- Ducat DC, Way JC, Silver PA. Engineering cyanobacteria to generate high-value products. Trends Biotechnol. 2011;29(2):95–103. - PubMed
-
- Zhang S, Bryant DA. The tricarboxylic acid cycle in cyanobacteria. Science. 2011;334(6062):1551–1553. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

