Ribosomal Proteins Rpl22 and Rpl22l1 Control Morphogenesis by Regulating Pre-mRNA Splicing
- PMID: 28076796
- PMCID: PMC5234864
- DOI: 10.1016/j.celrep.2016.12.034
Ribosomal Proteins Rpl22 and Rpl22l1 Control Morphogenesis by Regulating Pre-mRNA Splicing
Abstract
Most ribosomal proteins (RP) are regarded as essential, static components that contribute only to ribosome biogenesis and protein synthesis. However, emerging evidence suggests that RNA-binding RP are dynamic and can influence cellular processes by performing "extraribosomal," regulatory functions involving binding to select critical target mRNAs. We report here that the RP, Rpl22, and its highly homologous paralog Rpl22-Like1 (Rpl22l1 or Like1) play critical, extraribosomal roles in embryogenesis. Indeed, they antagonistically control morphogenesis through developmentally regulated localization to the nucleus, where they modulate splicing of the pre-mRNA encoding smad2, an essential transcriptional effector of Nodal/TGF-β signaling. During gastrulation, Rpl22 binds to intronic sequences of smad2 pre-mRNA and induces exon 9 skipping in cooperation with hnRNP-A1. This action is opposed by its paralog, Like1, which promotes exon 9 inclusion in the mature transcript. The nuclear roles of these RP in controlling morphogenesis represent a fundamentally different and paradigm-shifting mode of action for RP.
Keywords: Rpl22; Rpl22l1; Smad2; extraribosomal function; gastrulation; hnRNP-A1; morphogenesis; paralog; pre-mRNA splicing; ribosomal protein.
Copyright © 2017 The Author(s). Published by Elsevier Inc. All rights reserved.
Figures





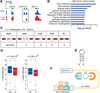

References
-
- Anderson SJ, Lauritsen JP, Hartman MG, Foushee AM, Lefebvre JM, Shinton SA, Gerhardt B, Hardy RR, Oravecz T, Wiest DL. Ablation of ribosomal protein L22 selectively impairs alphabeta T cell development by activation of a p53-dependent checkpoint. Immunity. 2007;26:759–772. - PubMed
-
- Dick A, Mayr T, Bauer H, Meier A, Hammerschmidt M. Cloning and characterization of zebrafish smad2, smad3 and smad4. Gene. 2000;246:69–80. - PubMed
-
- Dolnik A, Engelmann JC, Scharfenberger-Schmeer M, Mauch J, Kelkenberg-Schade S, Haldemann B, Fries T, Kronke J, Kuhn MW, Paschka P, et al. Commonly altered genomic regions in acute myeloid leukemia are enriched for somatic mutations involved in chromatin remodeling and splicing. Blood. 2012;120:e83–e92. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

