Optimizing Training Population Size and Genotyping Strategy for Genomic Prediction Using Association Study Results and Pedigree Information. A Case of Study in Advanced Wheat Breeding Lines
- PMID: 28081208
- PMCID: PMC5231327
- DOI: 10.1371/journal.pone.0169606
Optimizing Training Population Size and Genotyping Strategy for Genomic Prediction Using Association Study Results and Pedigree Information. A Case of Study in Advanced Wheat Breeding Lines
Abstract
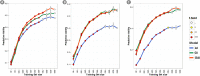
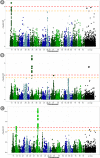
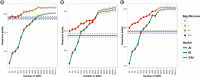
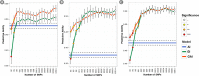
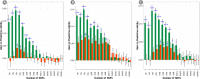
Wheat breeding programs generate a large amount of variation which cannot be completely explored because of limited phenotyping throughput. Genomic prediction (GP) has been proposed as a new tool which provides breeding values estimations without the need of phenotyping all the material produced but only a subset of it named training population (TP). However, genotyping of all the accessions under analysis is needed and, therefore, optimizing TP dimension and genotyping strategy is pivotal to implement GP in commercial breeding schemes. Here, we explored the optimum TP size and we integrated pedigree records and genome wide association studies (GWAS) results to optimize the genotyping strategy. A total of 988 advanced wheat breeding lines were genotyped with the Illumina 15K SNPs wheat chip and phenotyped across several years and locations for yield, lodging, and starch content. Cross-validation using the largest possible TP size and all the SNPs available after editing (~11k), yielded predictive abilities (rGP) ranging between 0.5-0.6. In order to explore the Training population size, rGP were computed using progressively smaller TP. These exercises showed that TP of around 700 lines were enough to yield the highest observed rGP. Moreover, rGP were calculated by randomly reducing the SNPs number. This showed that around 1K markers were enough to reach the highest observed rGP. GWAS was used to identify markers associated with the traits analyzed. A GWAS-based selection of SNPs resulted in increased rGP when compared with random selection and few hundreds SNPs were sufficient to obtain the highest observed rGP. For each of these scenarios, advantages of adding the pedigree information were shown. Our results indicate that moderate TP sizes were enough to yield high rGP and that pedigree information and GWAS results can be used to greatly optimize the genotyping strategy.
Conflict of interest statement
The authors have declared that no competing interests exist. This research was partly funded by the commercial partner Nordic seed A/S. This does not alter our adherence to PLOS ONE policies on sharing data and materials.
Figures
References
-
- Bernardo R. Quantitative traits in plants [Internet]. II edition Stemma press; 2010. Available: http://stemmapress.com
-
- Xu Y, Crouch JH. Marker-assisted selection in plant breeding: From publications to practice. Crop Sci. 2008;48: 391–407.
-
- Bernardo R. Molecular markers and selection for complex traits in plants: Learning from the last 20 years. Crop Sci. 2008;48: 1649–1664.
-
- Heffner EL, Sorrells ME, Jannink J-L. Genomic Selection for Crop Improvement. Crop Sci. 2009;49: 1.
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

