PDX1 dynamically regulates pancreatic ductal adenocarcinoma initiation and maintenance
- PMID: 28087712
- PMCID: PMC5238727
- DOI: 10.1101/gad.291021.116
PDX1 dynamically regulates pancreatic ductal adenocarcinoma initiation and maintenance
Abstract
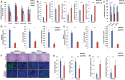
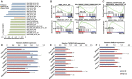
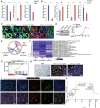
Aberrant activation of embryonic signaling pathways is frequent in pancreatic ductal adenocarcinoma (PDA), making developmental regulators therapeutically attractive. Here we demonstrate diverse functions for pancreatic and duodenal homeobox 1 (PDX1), a transcription factor indispensable for pancreas development, in the progression from normal exocrine cells to metastatic PDA. We identify a critical role for PDX1 in maintaining acinar cell identity, thus resisting the formation of pancreatic intraepithelial neoplasia (PanIN)-derived PDA. Upon neoplastic transformation, the role of PDX1 changes from tumor-suppressive to oncogenic. Interestingly, subsets of malignant cells lose PDX1 expression while undergoing epithelial-to-mesenchymal transition (EMT), and PDX1 loss is associated with poor outcome. This stage-specific functionality arises from profound shifts in PDX1 chromatin occupancy from acinar cells to PDA. In summary, we report distinct roles of PDX1 at different stages of PDA, suggesting that therapeutic approaches against this potential target need to account for its changing functions at different stages of carcinogenesis. These findings provide insight into the complexity of PDA pathogenesis and advocate a rigorous investigation of therapeutically tractable targets at distinct phases of PDA development and progression.
Keywords: EMT; dedifferentiation; pancreatic cancer; pancreatitis.
© 2016 Roy et al.; Published by Cold Spring Harbor Laboratory Press.
Figures




Similar articles
-
Pdx1 expression in pancreatic precursor lesions and neoplasms.Appl Immunohistochem Mol Morphol. 2011 Oct;19(5):444-9. doi: 10.1097/PAI.0b013e318206d958. Appl Immunohistochem Mol Morphol. 2011. PMID: 21297446 Free PMC article.
-
Brg1 promotes both tumor-suppressive and oncogenic activities at distinct stages of pancreatic cancer formation.Genes Dev. 2015 Mar 15;29(6):658-71. doi: 10.1101/gad.256628.114. Genes Dev. 2015. PMID: 25792600 Free PMC article.
-
The BRG1/SOX9 axis is critical for acinar cell-derived pancreatic tumorigenesis.J Clin Invest. 2018 Aug 1;128(8):3475-3489. doi: 10.1172/JCI94287. Epub 2018 Jul 16. J Clin Invest. 2018. PMID: 30010625 Free PMC article.
-
Transition to pancreatic cancer in response to carcinogen.Langenbecks Arch Surg. 2008 Jul;393(4):557-60. doi: 10.1007/s00423-007-0274-2. Epub 2008 Jan 12. Langenbecks Arch Surg. 2008. PMID: 18189145 Review.
-
PDX1: A Unique Pancreatic Master Regulator Constantly Changes Its Functions during Embryonic Development and Progression of Pancreatic Cancer.Biochemistry (Mosc). 2017 Aug;82(8):887-893. doi: 10.1134/S000629791708003X. Biochemistry (Mosc). 2017. PMID: 28941456 Review.
Cited by
-
Unveiling the Intricacies of Monoamine Oxidase-A (MAO-A) Inhibition in Colorectal Cancer: Computational Systems Biology, Expression Patterns, and the Anticancer Therapeutic Potential.ACS Omega. 2024 Aug 9;9(33):35703-35717. doi: 10.1021/acsomega.4c04100. eCollection 2024 Aug 20. ACS Omega. 2024. PMID: 39184489 Free PMC article.
-
Regulome-wide association study identifies genetically driven accessible regions associated with pancreatic cancer risk.Int J Cancer. 2024 Feb 15;154(4):670-678. doi: 10.1002/ijc.34761. Epub 2023 Oct 18. Int J Cancer. 2024. PMID: 37850323 Free PMC article.
-
Loss of activating transcription factor 3 prevents KRAS-mediated pancreatic cancer.Oncogene. 2021 Apr;40(17):3118-3135. doi: 10.1038/s41388-021-01771-z. Epub 2021 Apr 16. Oncogene. 2021. PMID: 33864001 Free PMC article.
-
Analysis of circulating blood and tissue biopsy PDX1 and MSX2 gene expression in patients with pancreatic cancer: A case-control experimental study.Medicine (Baltimore). 2019 Jun;98(26):e15954. doi: 10.1097/MD.0000000000015954. Medicine (Baltimore). 2019. PMID: 31261501 Free PMC article.
-
Pancreatic acinar cell fate relies on system xC- to prevent ferroptosis during stress.Cell Death Dis. 2023 Aug 21;14(8):536. doi: 10.1038/s41419-023-06063-w. Cell Death Dis. 2023. PMID: 37604805 Free PMC article.
References
-
- Bailey P, Chang DK, Nones K, Johns AL, Patch A-M, Gingras M-C, Miller DK, Christ AN, Bruxner TJC, Quinn MC, et al. 2016. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 531: 47–52. - PubMed
-
- Chen L, Yan H-X, Chen J, Yang W, Liu Q, Zhai B, Cao H-F, Liu S-Q, Wu M-C, Wang H-Y. 2007. Negative regulation of c-Myc transcription by pancreas duodenum homeobox-1. Endocrinology 148: 2168–2180. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
