Control of Amino Acid Homeostasis by a Ubiquitin Ligase-Coactivator Protein Complex
- PMID: 28100770
- PMCID: PMC5339764
- DOI: 10.1074/jbc.M116.766469
Control of Amino Acid Homeostasis by a Ubiquitin Ligase-Coactivator Protein Complex
Abstract
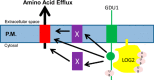
Intercellular amino acid transport is essential for the growth of all multicellular organisms, and its dysregulation is implicated in developmental disorders. By an unknown mechanism, amino acid efflux is stimulated in plants by overexpression of a membrane-localized protein (
Keywords: E3 ubiquitin ligase; Loss of GDU1 2; amino acid resistance; amino acid transport; glutamine dumper; protein degradation; ubiquitin; ubiquitin ligase; ubiquitylation (ubiquitination).
© 2017 by The American Society for Biochemistry and Molecular Biology, Inc.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article
Figures



Similar articles
-
The ubiquitin E3 ligase LOSS OF GDU2 is required for GLUTAMINE DUMPER1-induced amino acid secretion in Arabidopsis.Plant Physiol. 2012 Apr;158(4):1628-42. doi: 10.1104/pp.111.191965. Epub 2012 Jan 30. Plant Physiol. 2012. PMID: 22291198 Free PMC article.
-
Functional conservation between mammalian MGRN1 and plant LOG2 ubiquitin ligases.FEBS Lett. 2013 Nov 1;587(21):3400-5. doi: 10.1016/j.febslet.2013.08.045. Epub 2013 Sep 10. FEBS Lett. 2013. PMID: 24036454 Free PMC article.
-
The Arabidopsis RING E3 ubiquitin ligase AtAIRP3/LOG2 participates in positive regulation of high-salt and drought stress responses.Plant Physiol. 2013 Jul;162(3):1733-49. doi: 10.1104/pp.113.220103. Epub 2013 May 21. Plant Physiol. 2013. PMID: 23696092 Free PMC article.
-
Suppressor mutations in the Glutamine Dumper1 protein dissociate disturbance in amino acid transport from other characteristics of the Gdu1D phenotype.Front Plant Sci. 2015 Aug 4;6:593. doi: 10.3389/fpls.2015.00593. eCollection 2015. Front Plant Sci. 2015. PMID: 26300894 Free PMC article.
-
Ubiquitylation of ABA Receptors and Protein Phosphatase 2C Coreceptors to Modulate ABA Signaling and Stress Response.Int J Mol Sci. 2021 Jul 1;22(13):7103. doi: 10.3390/ijms22137103. Int J Mol Sci. 2021. PMID: 34281157 Free PMC article. Review.
Cited by
-
Identification and biochemical characterization of the fructokinase gene family in Arabidopsis thaliana.BMC Plant Biol. 2017 Apr 26;17(1):83. doi: 10.1186/s12870-017-1031-5. BMC Plant Biol. 2017. PMID: 28441933 Free PMC article.
-
The UBC27-AIRP3 ubiquitination complex modulates ABA signaling by promoting the degradation of ABI1 in Arabidopsis.Proc Natl Acad Sci U S A. 2020 Nov 3;117(44):27694-27702. doi: 10.1073/pnas.2007366117. Epub 2020 Oct 19. Proc Natl Acad Sci U S A. 2020. PMID: 33077597 Free PMC article.
-
Receptor control by membrane-tethered ubiquitin ligases in development and tissue homeostasis.Curr Top Dev Biol. 2022;150:25-89. doi: 10.1016/bs.ctdb.2022.03.003. Epub 2022 Jun 11. Curr Top Dev Biol. 2022. PMID: 35817504 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

