Mid-day siesta in natural populations of D. melanogaster from Africa exhibits an altitudinal cline and is regulated by splicing of a thermosensitive intron in the period clock gene
- PMID: 28114910
- PMCID: PMC5259850
- DOI: 10.1186/s12862-017-0880-8
Mid-day siesta in natural populations of D. melanogaster from Africa exhibits an altitudinal cline and is regulated by splicing of a thermosensitive intron in the period clock gene
Abstract
Background: Many diurnal animals exhibit a mid-day 'siesta', generally thought to be an adaptive response aimed at minimizing exposure to heat on warm days, suggesting that in regions with cooler climates mid-day siestas might be a less prominent feature of animal behavior. Drosophila melanogaster exhibits thermal plasticity in its mid-day siesta that is partly governed by the thermosensitive splicing of the 3'-terminal intron (termed dmpi8) from the key circadian clock gene period (per). For example, decreases in temperature lead to progressively more efficient splicing, which increasingly favors activity over sleep during the mid-day. In this study we sought to determine if the adaptation of D. melanogaster from its ancestral range in the lowlands of tropical Africa to the cooler temperatures found at high altitudes involved changes in mid-day sleep behavior and/or dmpi8 splicing efficiency.
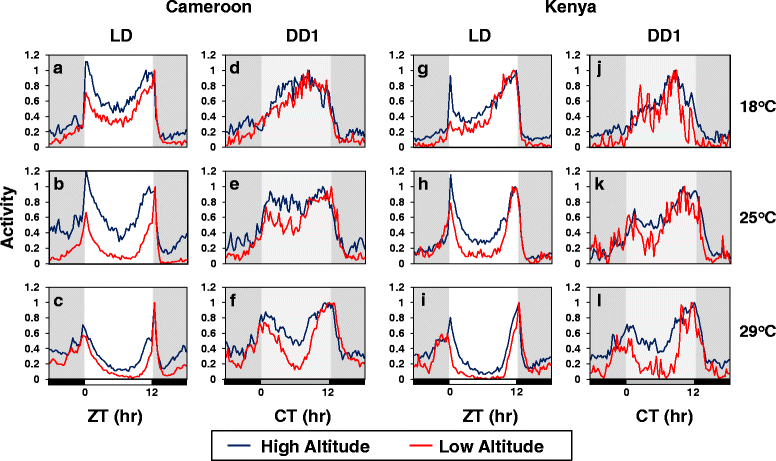
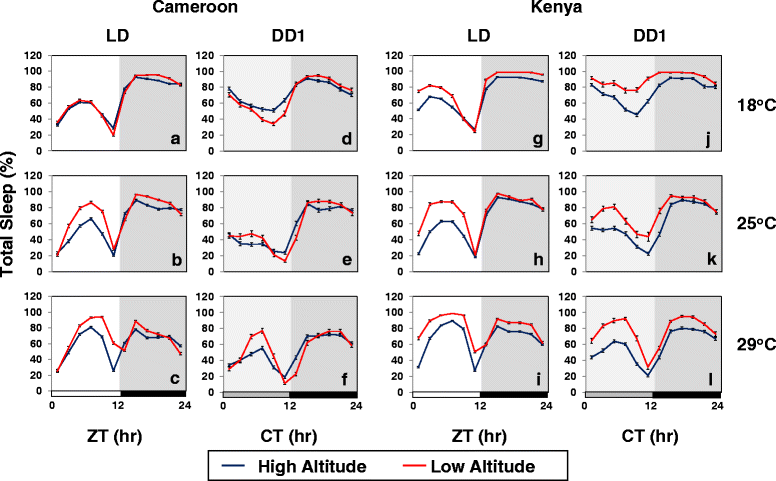
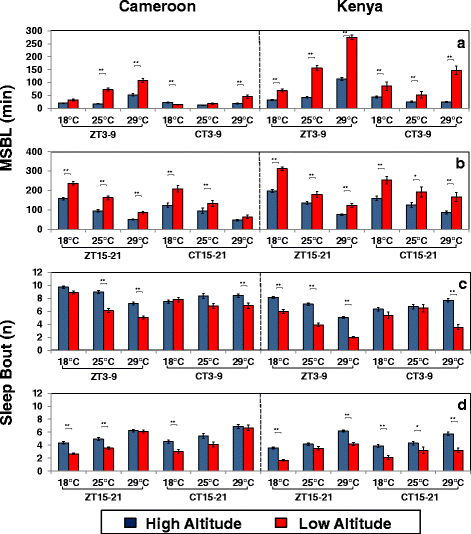
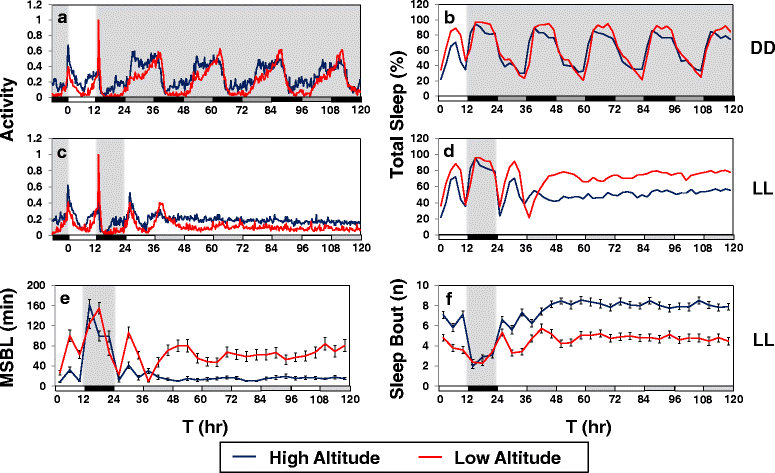
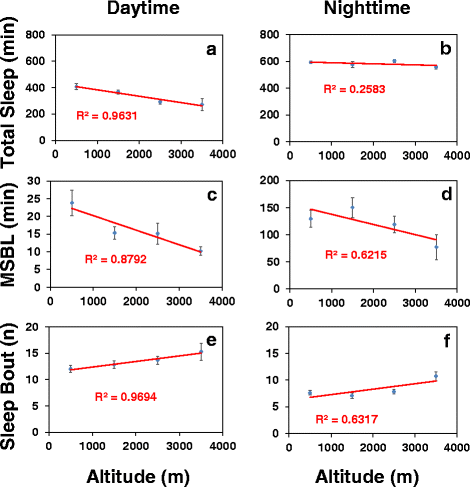
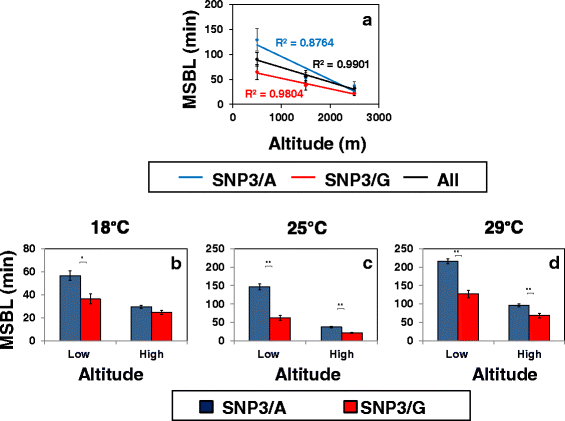
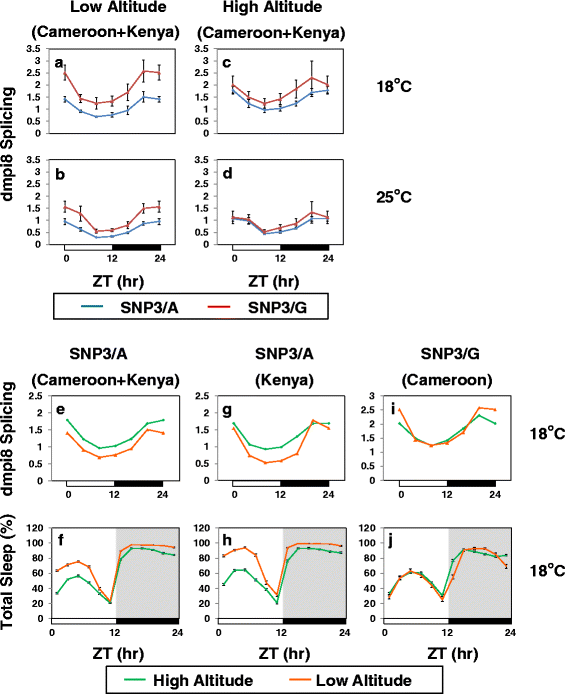
Results: Using natural populations of Drosophila melanogaster from different altitudes in tropical Africa we show that flies from high elevations have a reduced mid-day siesta and less consolidated sleep. We identified a single nucleotide polymorphism (SNP) in the per 3' untranslated region that has strong effects on dmpi8 splicing and mid-day sleep levels in both low and high altitude flies. Intriguingly, high altitude flies with a particular variant of this SNP exhibit increased dmpi8 splicing efficiency compared to their low altitude counterparts, consistent with reduced mid-day siesta. Thus, a boost in dmpi8 splicing efficiency appears to have played a prominent but not universal role in how African flies adapted to the cooler temperatures at high altitude.
Conclusions: Our findings point towards mid-day sleep behavior as a key evolutionary target in the thermal adaptation of animals, and provide a genetic framework for investigating daytime sleep in diurnal animals which appears to be driven by mechanisms distinct from those underlying nighttime sleep.
Keywords: Altitude; Circadian; Drosophila; Mid-day siesta; Period gene; Sleep; Splicing; Temperature; Thermal adaptation; dmpi8 intron.
Figures
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

