A Lin28 homologue reprograms differentiated cells to stem cells in the moss Physcomitrella patens
- PMID: 28128346
- PMCID: PMC5290140
- DOI: 10.1038/ncomms14242
A Lin28 homologue reprograms differentiated cells to stem cells in the moss Physcomitrella patens
Abstract
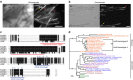
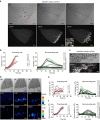
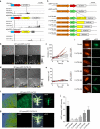
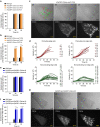
Both land plants and metazoa have the capacity to reprogram differentiated cells to stem cells. Here we show that the moss Physcomitrella patens Cold-Shock Domain Protein 1 (PpCSP1) regulates reprogramming of differentiated leaf cells to chloronema apical stem cells and shares conserved domains with the induced pluripotent stem cell factor Lin28 in mammals. PpCSP1 accumulates in the reprogramming cells and is maintained throughout the reprogramming process and in the resultant stem cells. Expression of PpCSP1 is negatively regulated by its 3'-untranslated region (3'-UTR). Removal of the 3'-UTR stabilizes PpCSP1 transcripts, results in accumulation of PpCSP1 protein and enhances reprogramming. A quadruple deletion mutant of PpCSP1 and three closely related PpCSP genes exhibits attenuated reprogramming indicating that the PpCSP genes function redundantly in cellular reprogramming. Taken together, these data demonstrate a positive role of PpCSP1 in reprogramming, which is similar to the function of mammalian Lin28.
Conflict of interest statement
The authors declare no competing financial interests.
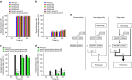
Figures
Similar articles
-
WOX13-like genes are required for reprogramming of leaf and protoplast cells into stem cells in the moss Physcomitrella patens.Development. 2014 Apr;141(8):1660-70. doi: 10.1242/dev.097444. Development. 2014. PMID: 24715456
-
Cells reprogramming to stem cells inhibit the reprogramming of adjacent cells in the moss Physcomitrella patens.Sci Rep. 2017 May 15;7(1):1909. doi: 10.1038/s41598-017-01786-1. Sci Rep. 2017. PMID: 28507289 Free PMC article.
-
Physcomitrella cyclin-dependent kinase A links cell cycle reactivation to other cellular changes during reprogramming of leaf cells.Plant Cell. 2011 Aug;23(8):2924-38. doi: 10.1105/tpc.111.088005. Epub 2011 Aug 23. Plant Cell. 2011. PMID: 21862705 Free PMC article.
-
Molecular mechanisms of reprogramming of differentiated cells into stem cells in the moss Physcomitrium patens.Curr Opin Plant Biol. 2022 Feb;65:102123. doi: 10.1016/j.pbi.2021.102123. Epub 2021 Nov 1. Curr Opin Plant Biol. 2022. PMID: 34735974 Review.
-
Cell cycle reentry from the late S phase: implications from stem cell formation in the moss Physcomitrella patens.J Plant Res. 2015 May;128(3):399-405. doi: 10.1007/s10265-015-0713-z. Epub 2015 Mar 24. J Plant Res. 2015. PMID: 25801272 Review.
Cited by
-
Central nervous system regeneration.Cell. 2022 Jan 6;185(1):77-94. doi: 10.1016/j.cell.2021.10.029. Cell. 2022. PMID: 34995518 Free PMC article. Review.
-
Digital Holographic Multimodal Cross-Sectional Fluorescence and Quantitative Phase Imaging System.Sci Rep. 2020 May 15;10(1):7580. doi: 10.1038/s41598-020-64028-x. Sci Rep. 2020. PMID: 32415184 Free PMC article.
-
LIN28A: A multifunctional versatile molecule with future therapeutic potential.World J Biol Chem. 2022 Mar 27;13(2):35-46. doi: 10.4331/wjbc.v13.i2.35. World J Biol Chem. 2022. PMID: 35432768 Free PMC article. Review.
-
Lin28 Signaling Supports Mammalian PNS and CNS Axon Regeneration.Cell Rep. 2018 Sep 4;24(10):2540-2552.e6. doi: 10.1016/j.celrep.2018.07.105. Cell Rep. 2018. PMID: 30184489 Free PMC article.
-
Co-option of plant gene regulatory network in nutrient responses during terrestrialization.Nat Plants. 2024 Dec;10(12):1955-1968. doi: 10.1038/s41477-024-01851-4. Epub 2024 Nov 26. Nat Plants. 2024. PMID: 39592744
References
-
- Heidstra R. & Sabatini S. Plant and animal stem cells: similar yet different. Nat. Rev. Mol. Cell Biol. 15, 301–312 (2014). - PubMed
-
- Sugimoto K., Gordon S. P. & Meyerowitz E. M. Regeneration in plants and animals: dedifferentiation, transdifferentiation, or just differentiation? Trends Cell Biol. 21, 212–218 (2011). - PubMed
-
- Steward F., Mapes M. O. & Mears K. Growth and organized development of cultured cells. II. Organization in cultures grown from freely suspended cells. Am. J. Bot. 45, 705–708 (1958).
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

