Studying Brain Circuit Function with Dynamic Causal Modeling for Optogenetic fMRI
- PMID: 28132829
- PMCID: PMC5472443
- DOI: 10.1016/j.neuron.2016.12.035
Studying Brain Circuit Function with Dynamic Causal Modeling for Optogenetic fMRI
Abstract
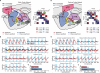
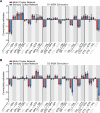
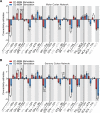
Defining the large-scale behavior of brain circuits with cell type specificity is a major goal of neuroscience. However, neuronal circuit diagrams typically draw upon anatomical and electrophysiological measurements acquired in isolation. Consequently, a dynamic and cell-type-specific connectivity map has never been constructed from simultaneous measurements across the brain. Here, we introduce dynamic causal modeling (DCM) for optogenetic fMRI experiments-which uniquely allow cell-type-specific, brain-wide functional measurements-to parameterize the causal relationships among regions of a distributed brain network with cell type specificity. Strikingly, when applied to the brain-wide basal ganglia-thalamocortical network, DCM accurately reproduced the empirically observed time series, and the strongest connections were key connections of optogenetically stimulated pathways. We predict that quantitative and cell-type-specific descriptions of dynamic connectivity, as illustrated here, will empower novel systems-level understanding of neuronal circuit dynamics and facilitate the design of more effective neuromodulation therapies.
Keywords: basal ganglia; direct and indirect pathways; dynamic causal modeling; ofMRI; optogenetic fMRI.
Copyright © 2017 Elsevier Inc. All rights reserved.
Figures

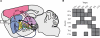

References
-
- Abe Y, Sekino M, Terazono Y, Ohsaki H, Fukazawa Y, Sakai S, Yawo H, Hisatsune T. Opto-fMRI analysis for exploring the neuronal connectivity of the hippocampal formation in rats. Neuroscience research. 2012;74:248–255. - PubMed
-
- Albin RL, Young AB, Penney JB. The functional anatomy of basal ganglia disorders. Trends in neurosciences. 1989;12:366–375. - PubMed
-
- Bandettini PA. Twenty years of functional MRI: the science and the stories. Neuroimage. 2012;62:575–588. - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

