Spheroid arrays for high-throughput single-cell analysis of spatial patterns and biomarker expression in 3D
- PMID: 28134245
- PMCID: PMC5278392
- DOI: 10.1038/srep41160
Spheroid arrays for high-throughput single-cell analysis of spatial patterns and biomarker expression in 3D
Abstract
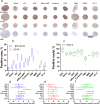
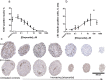
We describe and share a device, methodology and image analysis algorithms, which allow up to 66 spheroids to be arranged into a gel-based array directly from a culture plate for downstream processing and analysis. Compared to processing individual samples, the technique uses 11-fold less reagents, saves time and enables automated imaging. To illustrate the power of the technology, we showcase applications of the methodology for investigating 3D spheroid morphology and marker expression and for in vitro safety and efficacy screens. First, spheroid arrays of 11 cell-lines were rapidly assessed for differences in spheroid morphology. Second, highly-positive (SOX-2), moderately-positive (Ki-67) and weakly-positive (βIII-tubulin) protein targets were detected and quantified. Third, the arrays enabled screening of ten media compositions for inducing differentiation in human neurospheres. Last, the application of spheroid microarrays for spheroid-based drug screens was demonstrated by quantifying the dose-dependent drop in proliferation and increase in differentiation in etoposide-treated neurospheres.
Conflict of interest statement
The authors declare no competing financial interests.
Figures




References
-
- Olson H. et al.. Concordance of the toxicity of pharmaceuticals in humans and in animals. Regul. Toxicol. Pharmacol. 32, 56–67 (2000). - PubMed
-
- Hickman J. A. et al.. Three-dimensional models of cancer for pharmacology and cancer cell biology: capturing tumor complexity in vitro/ex vivo. Biotechnol. J. 9, 1115–28 (2014). - PubMed
-
- Sutherland R. M., McCredie J. a. & Inch W. R. Growth of multicell spheroids in tissue culture as a model of nodular carcinomas. J. Natl. Cancer Inst. 46, 113–20 (1971). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

