Cell-to-cell heterogeneity of EWSR1-FLI1 activity determines proliferation/migration choices in Ewing sarcoma cells
- PMID: 28135250
- PMCID: PMC5541267
- DOI: 10.1038/onc.2016.498
Cell-to-cell heterogeneity of EWSR1-FLI1 activity determines proliferation/migration choices in Ewing sarcoma cells
Abstract
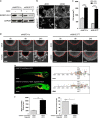
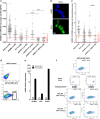
Ewing sarcoma is characterized by the expression of the chimeric EWSR1-FLI1 transcription factor. Proteomic analyses indicate that the decrease of EWSR1-FLI1 expression leads to major changes in effectors of the dynamics of the actin cytoskeleton and the adhesion processes with a shift from cell-to-cell to cell-matrix adhesion. These changes are associated with a dramatic increase of in vivo cell migration and invasion potential. Importantly, EWSR1-FLI1 expression, evaluated by single-cell RT-ddPCR/immunofluorescence analyses, and activity, assessed by expression of EWSR1-FLI1 downstream targets, are heterogeneous in cell lines and in tumours and can fluctuate along time in a fully reversible process between EWSR1-FLI1high states, characterized by highly active cell proliferation, and EWSR1-FLI1low states where cells have a strong propensity to migrate, invade and metastasize. This new model of phenotypic plasticity proposes that the dynamic fluctuation of the expression level of a dominant oncogene is an intrinsic characteristic of its oncogenic potential.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- HaDuong JH, Martin AA, Skapek SX, Mascarenhas L. Sarcomas. Pediatr Clin North Am 2015; 62: 179–200. - PubMed
-
- Deley M-CL, Paulussen M, Lewis I, Brennan B, Ranft A, Whelan J et al. Cyclophosphamide compared with ifosfamide in consolidation treatment of standard-risk Ewing sarcoma: results of the randomized noninferiority euro-EWING99-R1 trial. J Clin Oncol 2014; 32: 2440–2448. - PubMed
-
- Delattre O, Zucman J, Plougastel B, Desmaze C, Melot T, Peter M et al. Gene fusion with an ETS DNA-binding domain caused by chromosome translocation in human tumours. Nature 1992; 359: 162–165. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

