Transcription and replication mechanisms of Bunyaviridae and Arenaviridae L proteins
- PMID: 28137457
- PMCID: PMC7114536
- DOI: 10.1016/j.virusres.2017.01.018
Transcription and replication mechanisms of Bunyaviridae and Arenaviridae L proteins
Abstract
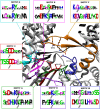
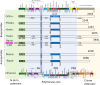
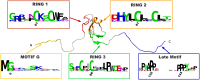
Bunyaviridae and Arenaviridae virus families include an important number of highly pathogenic viruses for humans. They are enveloped viruses with negative stranded RNA genomes divided into three (bunyaviruses) or two (arenaviruses) segments. Each genome segment is coated by the viral nucleoproteins (NPs) and the polymerase (L protein) to form a functional ribonucleoprotein (RNP) complex. The viral RNP provides the necessary context on which the L protein carries out the biosynthetic processes of RNA replication and gene transcription. Decades of research have provided a good understanding of the molecular processes underlying RNA synthesis, both RNA replication and gene transcription, for these two families of viruses. In this review we will provide a global view of the common features, as well as differences, of the molecular biology of Bunyaviridae and Arenaviridae. We will also describe structures of protein and protein-RNA complexes so far determined for these viral families, mainly focusing on the L protein, and discuss their implications for understanding the mechanisms of viral RNA replication and gene transcription within the architecture of viral RNPs, also taking into account the cellular context in which these processes occur. Finally, we will discuss the implications of these structural findings for the development of antiviral drugs to treat human diseases caused by members of the Bunyaviridae and Arenaviridae families.
Keywords: Antivirals; Arenavirus; Bunyavirus; L protein; Replication; Transcription.
Copyright © 2017 Elsevier B.V. All rights reserved.
Figures




References
-
- Andrei G., De Clercq E. Molecular approaches for the treatment of hemorrhagic fever virus infections. Antivir. Res. 1993;22:45–75. - PubMed
-
- Barber D.L., Wherry E.J., Masopust D., Zhu B., Allison J.P., Sharpe A.H., Freeman G.J., Ahmed R. Restoring function in exhausted CD8 T cells during chronic viral infection. Nature. 2006;439:682–687. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

