RNA-Puzzles Round III: 3D RNA structure prediction of five riboswitches and one ribozyme
- PMID: 28138060
- PMCID: PMC5393176
- DOI: 10.1261/rna.060368.116
RNA-Puzzles Round III: 3D RNA structure prediction of five riboswitches and one ribozyme
Abstract
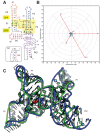
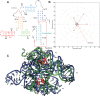
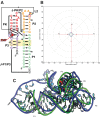
RNA-Puzzles is a collective experiment in blind 3D RNA structure prediction. We report here a third round of RNA-Puzzles. Five puzzles, 4, 8, 12, 13, 14, all structures of riboswitch aptamers and puzzle 7, a ribozyme structure, are included in this round of the experiment. The riboswitch structures include biological binding sites for small molecules (S-adenosyl methionine, cyclic diadenosine monophosphate, 5-amino 4-imidazole carboxamide riboside 5'-triphosphate, glutamine) and proteins (YbxF), and one set describes large conformational changes between ligand-free and ligand-bound states. The Varkud satellite ribozyme is the most recently solved structure of a known large ribozyme. All puzzles have established biological functions and require structural understanding to appreciate their molecular mechanisms. Through the use of fast-track experimental data, including multidimensional chemical mapping, and accurate prediction of RNA secondary structure, a large portion of the contacts in 3D have been predicted correctly leading to similar topologies for the top ranking predictions. Template-based and homology-derived predictions could predict structures to particularly high accuracies. However, achieving biological insights from de novo prediction of RNA 3D structures still depends on the size and complexity of the RNA. Blind computational predictions of RNA structures already appear to provide useful structural information in many cases. Similar to the previous RNA-Puzzles Round II experiment, the prediction of non-Watson-Crick interactions and the observed high atomic clash scores reveal a notable need for an algorithm of improvement. All prediction models and assessment results are available at http://ahsoka.u-strasbg.fr/rnapuzzles/.
Keywords: 3D prediction; X-ray structures; bioinformatics; force fields; models; structure quality.
© 2017 Miao et al.; Published by Cold Spring Harbor Laboratory Press for the RNA Society.
Figures



References
-
- Antczak M, Popenda M, Zok T, Sarzynska J, Ratajczak T, Adamiak RW, Szachniuk M. 2016. New functionality of RNAComposer: application to shape the axis of miR160 precursor structure. Acta Biochim Pol 63: 737–744. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
