Transcriptional Profiling in Experimental Visceral Leishmaniasis Reveals a Broad Splenic Inflammatory Environment that Conditions Macrophages toward a Disease-Promoting Phenotype
- PMID: 28141856
- PMCID: PMC5283737
- DOI: 10.1371/journal.ppat.1006165
Transcriptional Profiling in Experimental Visceral Leishmaniasis Reveals a Broad Splenic Inflammatory Environment that Conditions Macrophages toward a Disease-Promoting Phenotype
Abstract
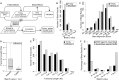
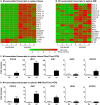
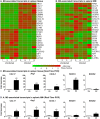
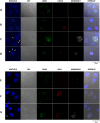
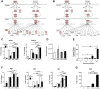
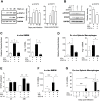
Visceral Leishmaniasis (VL), caused by the intracellular protozoan Leishmania donovani, is characterized by relentlessly increasing visceral parasite replication, cachexia, massive splenomegaly, pancytopenia and ultimately death. Progressive disease is considered to be due to impaired effector T cell function and/or failure of macrophages to be activated to kill the intracellular parasite. In previous studies, we used the Syrian hamster (Mesocricetus auratus) as a model because it mimics the progressive nature of active human VL. We demonstrated previously that mixed expression of macrophage-activating (IFN-γ) and regulatory (IL-4, IL-10, IL-21) cytokines, parasite-induced expression of macrophage arginase 1 (Arg1), and decreased production of nitric oxide are key immunopathologic factors. Here we examined global changes in gene expression to define the splenic environment and phenotype of splenic macrophages during progressive VL. We used RNA sequencing coupled with de novo transcriptome assembly, because the Syrian hamster does not have a fully sequenced and annotated reference genome. Differentially expressed transcripts identified a highly inflammatory spleen environment with abundant expression of type I and type II interferon response genes. However, high IFN-γ expression was ineffective in directing exclusive M1 macrophage polarization, suppressing M2-associated gene expression, and restraining parasite replication and disease. While many IFN-inducible transcripts were upregulated in the infected spleen, fewer were induced in splenic macrophages in VL. Paradoxically, IFN-γ enhanced parasite growth and induced the counter-regulatory molecules Arg1, Ido1 and Irg1 in splenic macrophages. This was mediated, at least in part, through IFN-γ-induced activation of STAT3 and expression of IL-10, which suggests that splenic macrophages in VL are conditioned to respond to macrophage activation signals with a counter-regulatory response that is ineffective and even disease-promoting. Accordingly, inhibition of STAT3 activation led to a reduced parasite load in infected macrophages. Thus, the STAT3 pathway offers a rational target for adjunctive host-directed therapy to interrupt the pathogenesis of VL.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures





Similar articles
-
Splenic CD4+ T Cells in Progressive Visceral Leishmaniasis Show a Mixed Effector-Regulatory Phenotype and Impair Macrophage Effector Function through Inhibitory Receptor Expression.PLoS One. 2017 Jan 19;12(1):e0169496. doi: 10.1371/journal.pone.0169496. eCollection 2017. PLoS One. 2017. PMID: 28103263 Free PMC article.
-
Transcriptional profiling of the spleen in progressive visceral leishmaniasis reveals mixed expression of type 1 and type 2 cytokine-responsive genes.BMC Immunol. 2014 Nov 26;15:38. doi: 10.1186/s12865-014-0038-z. BMC Immunol. 2014. PMID: 25424735 Free PMC article.
-
Progressive visceral leishmaniasis is driven by dominant parasite-induced STAT6 activation and STAT6-dependent host arginase 1 expression.PLoS Pathog. 2012 Jan;8(1):e1002417. doi: 10.1371/journal.ppat.1002417. Epub 2012 Jan 19. PLoS Pathog. 2012. PMID: 22275864 Free PMC article.
-
Balancing immunity and pathology in visceral leishmaniasis.Immunol Cell Biol. 2007 Feb-Mar;85(2):138-47. doi: 10.1038/sj.icb7100011. Epub 2006 Dec 5. Immunol Cell Biol. 2007. PMID: 17146466 Review.
-
Immunoactivation and immunopathogeny during active visceral leishmaniasis.Rev Inst Med Trop Sao Paulo. 2009 Sep-Oct;51(5):241-6. doi: 10.1590/s0036-46652009000500002. Rev Inst Med Trop Sao Paulo. 2009. PMID: 19893975 Review.
Cited by
-
Model-based Evaluation of Gene Expression Changes in Response to Leishmania Infection.Gene Regul Syst Bio. 2019 Feb 6;13:1177625019828350. doi: 10.1177/1177625019828350. eCollection 2019. Gene Regul Syst Bio. 2019. PMID: 30792575 Free PMC article. Review.
-
Splenic Transcriptional Responses in Severe Visceral Leishmaniasis: Impaired Leukocyte Chemotaxis and Cell Cycle Arrest.Front Immunol. 2021 Nov 5;12:716314. doi: 10.3389/fimmu.2021.716314. eCollection 2021. Front Immunol. 2021. PMID: 34804009 Free PMC article.
-
Integrated analysis of lncRNA and mRNA expression profiles in cutaneous leishmaniasis lesions caused by Leishmania tropica.Front Cell Infect Microbiol. 2024 Nov 21;14:1416925. doi: 10.3389/fcimb.2024.1416925. eCollection 2024. Front Cell Infect Microbiol. 2024. PMID: 39639867 Free PMC article.
-
Fatal progression of experimental visceral leishmaniasis is associated with intestinal parasitism and secondary infection by commensal bacteria, and is delayed by antibiotic prophylaxis.PLoS Pathog. 2020 Apr 13;16(4):e1008456. doi: 10.1371/journal.ppat.1008456. eCollection 2020 Apr. PLoS Pathog. 2020. PMID: 32282850 Free PMC article.
-
TIM-3 increases the abundance of type-2 dendritic cells during Leishmania donovani infection by enhancing IL-10 production via STAT3.Cell Death Dis. 2023 May 18;14(5):331. doi: 10.1038/s41419-023-05848-3. Cell Death Dis. 2023. PMID: 37202419 Free PMC article.
References
-
- World Health Organization (2014) Leishmaniasis: Situation and trends. Global Health Observatory (GHO) World Health Organization, http://www.who.int/gho/neglected_diseases/leishmaniasis/en/.
-
- Bacellar O, Brodskyn C, Guerreiro J, Barral-Netto M, Costa CH, et al. (1996) Interleukin-12 restores interferon-gamma production and cytotoxic responses in visceral leishmaniasis. J Infect Dis 173: 1515–1518. - PubMed
-
- Carvalho EM, Bacellar OA, Reed S, Barral A, Rocha H (1988) Visceral leishmaniasis: a disease associated with inability of lymphocytes to activate macrophages to kill leishmania. Braz J Med Biol Res 21: 85–92. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

