TP53 mutation-mediated genomic instability induces the evolution of chemoresistance and recurrence in epithelial ovarian cancer
- PMID: 28148293
- PMCID: PMC5288946
- DOI: 10.1186/s13000-017-0605-8
TP53 mutation-mediated genomic instability induces the evolution of chemoresistance and recurrence in epithelial ovarian cancer
Abstract
Background: Genomic instability caused by mutation of the checkpoint molecule TP53 may endow cancer cells with the ability to undergo genomic evolution to survive stress and treatment. We attempted to gain insight into the potential contribution of ovarian cancer genomic instability resulted from TP53 mutation to the aberrant expression of multidrug resistance gene MDR1.
Methods: TP53 mutation status was assessed by performing nucleotide sequencing and immunohistochemistry. Ovarian cancer cell DNA ploidy was determined using Feulgen-stained smears or flow cytometry. DNA copy number was analyzed by performing fluorescence in situ hybridization (FISH).
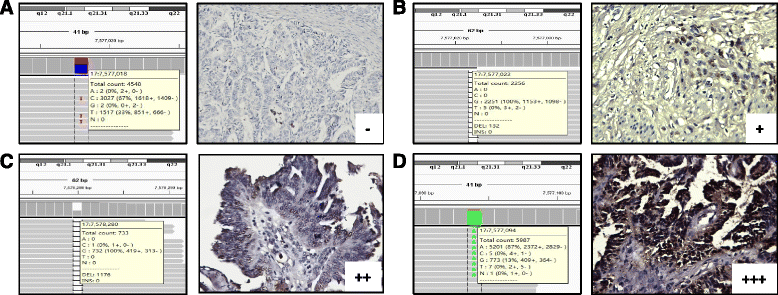
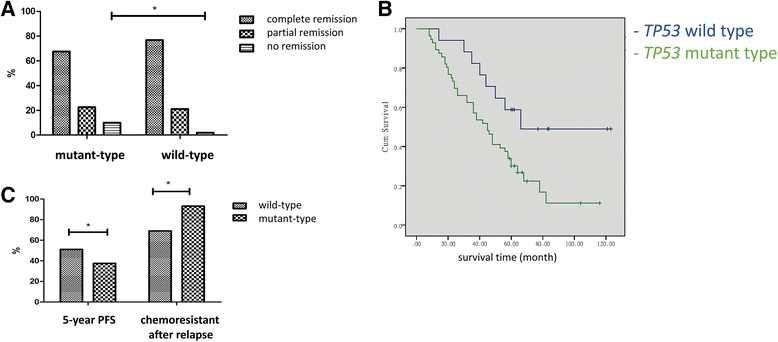
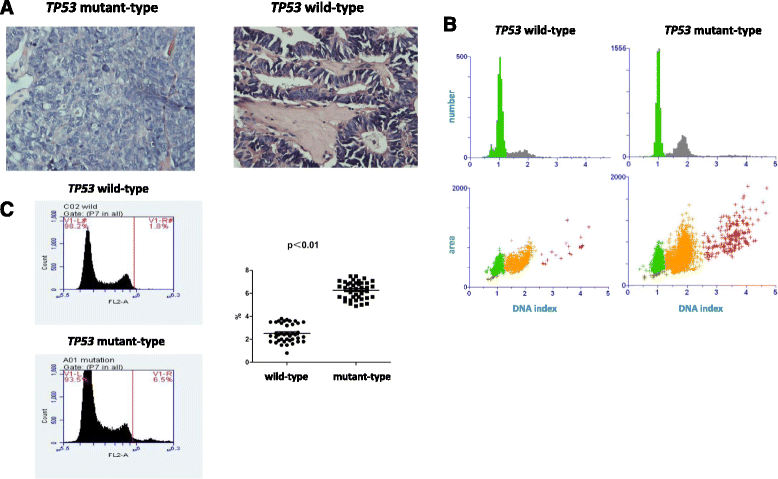
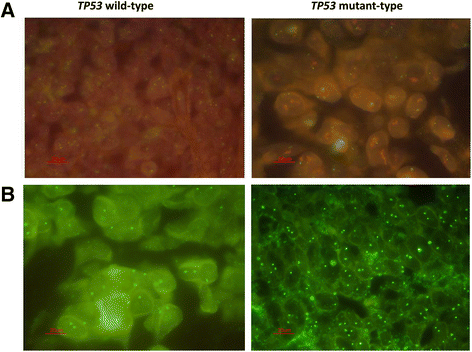
Results: In addition to performing nucleotide sequencing for 5 cases of ovarian cancer, TP53 mutations were analyzed via immunohistochemical staining for P53. Both intensive P53 immunohistochemical staining and complete absence of signal were associated with the occurrence of TP53 mutations. HE staining and the quantification of DNA content indicated a significantly higher proportion of polyploidy and aneuploidy cells in the TP53 mutant group than in the wild-type group (p < 0.05). Moreover, in 161 epithelial ovarian cancer patients, multivariate logistic analysis identified late FIGO (International Federation of Gynecology and Obstetrics) stage, serous histotype, G3 grade and TP53 mutation as independent risk factors for ovarian cancer recurrence. In relapse patients, the proportion of chemoresistant cases in the TP53 wild-type group was significantly lower than in the mutant group (63.6% vs. 91.8%, p < 0.05). FISH results revealed a higher percentage of cells with >6 MDR1 copies and chromosome 7 amplication in the TP53 mutant group than in the wild-type group [11.7 ± 2.3% vs. 3.0 ± 0.7% and 2.1 ± 0.7% vs. 0.3 ± 0.05%, (p < 0.05), respectively]. And we observed a specific increase of MDR1 and chromosome 7 copy numbers in the TP53 mutant group upon disease regression (p < 0.01).
Conclusions: TP53 mutation-associated genomic instability may promote chromosome 7 accumulation and MDR1 amplification during ovarian cancer chemoresistance and recurrence. Our findings lay the foundation for the development of promising chemotherapeutic approaches to treat aggressive and recurrent ovarian cancer.
Keywords: Epithelial ovarian cancer; Genomic instability; MDR1 copy number; Recurrence; TP53 mutation.
Figures

References
-
- Kovac M, Navas C, Horswell S, Salm M, Bardella C, Rowan A, Stares M, Castro-Giner F, Fisher R, de Bruin EC, Kovacova M, Gorman M, Makino S, Williams J, Jaeger E, Jones A, Howarth K, Larkin J, Pickering L, Gore M. Recurrent chromosomal gains and heterogeneous driver mutations characterise papillary renal cancer evolution. Nat Commun. 2015;6:6336. doi: 10.1038/ncomms7336. - DOI - PMC - PubMed
-
- Murugaesu N, Wilson GA, Birkbak NJ, Watkins TB, McGranahan N, Kumar S, Abbassi-Ghadi N, Salm M, Mitter R, Horswell S, Rowan A, Phillimore B, Biggs J, Begum S, Matthews N, Hochhauser D, Hanna GB, Swanton C. Tracking the genomic evolution of esophageal adenocarcinoma through neoadjuvant chemotherapy. Cancer Discov. 2015;5:821–31. doi: 10.1158/2159-8290.CD-15-0412. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

