The DNA methylation level against the background of the genome size and t-heterochromatin content in some species of the genus Secale L
- PMID: 28149679
- PMCID: PMC5267573
- DOI: 10.7717/peerj.2889
The DNA methylation level against the background of the genome size and t-heterochromatin content in some species of the genus Secale L
Abstract
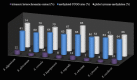
Methylation of cytosine in DNA is one of the most important epigenetic modifications in eukaryotes and plays a crucial role in the regulation of gene activity and the maintenance of genomic integrity. DNA methylation and other epigenetic mechanisms affect the development, differentiation or the response of plants to biotic and abiotic stress. This study compared the level of methylation of cytosines on a global (ELISA) and genomic scale (MSAP) between the species of the genus Secale. We analyzed whether the interspecific variation of cytosine methylation was associated with the size of the genome (C-value) and the content of telomeric heterochromatin. MSAP analysis showed that S. sylvestre was the most distinct species among the studied rye taxa; however, the results clearly indicated that these differences were not statistically significant. The total methylation level of the studied loci was very similar in all taxa and ranged from 60% in S. strictum ssp. africanum to 66% in S. cereale ssp. segetale, which confirmed the lack of significant differences in the sequence methylation pattern between the pairs of rye taxa. The level of global cytosine methylation in the DNA was not significantly associated with the content of t-heterochromatin and did not overlap with the existing taxonomic rye relationships. The highest content of 5-methylcytosine was found in S. cereale ssp. segetale (83%), while very low in S. strictum ssp. strictum (53%), which was significantly different from the methylation state of all taxa, except for S. sylvestre. The other studied taxa of rye had a similar level of methylated cytosine ranging from 66.42% (S. vavilovii) to 74.41% in (S. cereale ssp. afghanicum). The results obtained in this study are evidence that the percentage of methylated cytosine cannot be inferred solely based on the genome size or t-heterochromatin. This is a significantly more complex issue.
Keywords: ELISA; Genome size; MSAP; Methylation DNA; Secale; t-heterochromatin.
Conflict of interest statement
The authors declare there are no competing interests.
Figures





References
-
- Achrem M, Kalinka A, Rogalska S. Assessment of genetic relationships among Secale taxa by using ISSR and IRAP markers and the chromosomal distribution of the AAC microsatellite sequence. Turkish Journal of Botany. 2014;38:213–225. doi: 10.3906/bot-1207-26. - DOI
-
- Alonso C, Pérez R, Bazaga P, Medrano M, Herrera CM. Individual variation in size and fecundity is correlated with differences in global DNA cytosine methylation in the perennial herb Helleborus foetidus (Ranunculaceae) American Journal Botanical. 2014;101:1309–1313. doi: 10.3732/ajb.1400126. - DOI - PubMed
-
- Banks J, Fedoroff N. Patterns of developmental and heritable change in methylation of the suppressor-mutator transposable element. Developmental Genetics. 1989;10(6):425–437. doi: 10.1002/dvg.1020100604. - DOI
LinkOut - more resources
Full Text Sources
Other Literature Sources

