Dissecting the proton transport pathway in electrogenic Na+/H+ antiporters
- PMID: 28154142
- PMCID: PMC5321028
- DOI: 10.1073/pnas.1614521114
Dissecting the proton transport pathway in electrogenic Na+/H+ antiporters
Abstract
Sodium/proton exchangers of the SLC9 family mediate the transport of protons in exchange for sodium to help regulate intracellular pH, sodium levels, and cell volume. In electrogenic Na+/H+ antiporters, it has been assumed that two ion-binding aspartate residues transport the two protons that are later exchanged for one sodium ion. However, here we show that we can switch the antiport activity of the bacterial Na+/H+ antiporter NapA from being electrogenic to electroneutral by the mutation of a single lysine residue (K305). Electroneutral lysine mutants show similar ion affinities when driven by [Formula: see text]pH, but no longer respond to either an electrochemical potential ([Formula: see text]) or could generate one when driven by ion gradients. We further show that the exchange activity of the human Na+/H+ exchanger NHA2 (SLC9B2) is electroneutral, despite harboring the two conserved aspartic acid residues found in NapA and other bacterial homologues. Consistently, the equivalent residue to K305 in human NHA2 has been replaced with arginine, which is a mutation that makes NapA electroneutral. We conclude that a transmembrane embedded lysine residue is essential for electrogenic transport in Na+/H+ antiporters.
Keywords: Na+/H+exchangers; energetics; membrane protein; proton transport; secondary active transporters.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
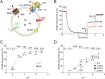
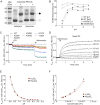


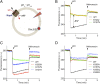



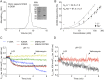
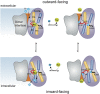
References
-
- Padan E. The enlightening encounter between structure and function in the NhaA Na+–H+ antiporter. Trends Biochem Sci. 2008;33(9):435–443. - PubMed
-
- Brett CL, Donowitz M, Rao R. Evolutionary origins of eukaryotic sodium/proton exchangers. Am J Physiol Cell Physiol. 2005;288(2):C223–C239. - PubMed
-
- Taglicht D, Padan E, Schuldiner S. Proton-sodium stoichiometry of NhaA, an electrogenic antiporter from Escherichia coli. J Biol Chem. 1993;268(8):5382–5387. - PubMed
-
- Pinner E, Padan E, Schuldiner S. Kinetic properties of NhaB, a Na+/H+ antiporter from Escherichia coli. J Biol Chem. 1994;269(42):26274–26279. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

