Reverse Methanogenesis and Respiration in Methanotrophic Archaea
- PMID: 28154498
- PMCID: PMC5244752
- DOI: 10.1155/2017/1654237
Reverse Methanogenesis and Respiration in Methanotrophic Archaea
Abstract
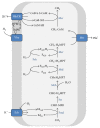
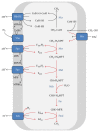
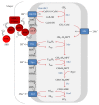
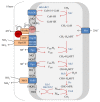
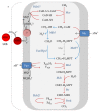
Anaerobic oxidation of methane (AOM) is catalyzed by anaerobic methane-oxidizing archaea (ANME) via a reverse and modified methanogenesis pathway. Methanogens can also reverse the methanogenesis pathway to oxidize methane, but only during net methane production (i.e., "trace methane oxidation"). In turn, ANME can produce methane, but only during net methane oxidation (i.e., enzymatic back flux). Net AOM is exergonic when coupled to an external electron acceptor such as sulfate (ANME-1, ANME-2abc, and ANME-3), nitrate (ANME-2d), or metal (oxides). In this review, the reversibility of the methanogenesis pathway and essential differences between ANME and methanogens are described by combining published information with domain based (meta)genome comparison of archaeal methanotrophs and selected archaea. These differences include abundances and special structure of methyl coenzyme M reductase and of multiheme cytochromes and the presence of menaquinones or methanophenazines. ANME-2a and ANME-2d can use electron acceptors other than sulfate or nitrate for AOM, respectively. Environmental studies suggest that ANME-2d are also involved in sulfate-dependent AOM. ANME-1 seem to use a different mechanism for disposal of electrons and possibly are less versatile in electron acceptors use than ANME-2. Future research will shed light on the molecular basis of reversal of the methanogenic pathway and electron transfer in different ANME types.
Conflict of interest statement
The authors declare that there is no conflict of interests regarding the publication of this paper.
Figures

Similar articles
-
Experimental evaluation of the metabolic reversibility of ANME-2d between anaerobic methane oxidation and methanogenesis.Appl Microbiol Biotechnol. 2016 Jul;100(14):6481-6490. doi: 10.1007/s00253-016-7475-y. Epub 2016 Mar 30. Appl Microbiol Biotechnol. 2016. PMID: 27026178
-
Anaerobic oxidation of methane coupled to nitrate reduction in a novel archaeal lineage.Nature. 2013 Aug 29;500(7464):567-70. doi: 10.1038/nature12375. Epub 2013 Jul 28. Nature. 2013. PMID: 23892779
-
Spatial-Temporal Pattern of Sulfate-Dependent Anaerobic Methane Oxidation in an Intertidal Zone of the East China Sea.Appl Environ Microbiol. 2019 Mar 22;85(7):e02638-18. doi: 10.1128/AEM.02638-18. Print 2019 Apr 1. Appl Environ Microbiol. 2019. PMID: 30709818 Free PMC article.
-
Anaerobic oxidation of methane: an "active" microbial process.Microbiologyopen. 2015 Feb;4(1):1-11. doi: 10.1002/mbo3.232. Epub 2014 Dec 22. Microbiologyopen. 2015. PMID: 25530008 Free PMC article. Review.
-
Metabolic potential of anaerobic methane oxidizing archaea for a broad spectrum of electron acceptors.Adv Microb Physiol. 2022;80:157-201. doi: 10.1016/bs.ampbs.2022.01.003. Epub 2022 Feb 18. Adv Microb Physiol. 2022. PMID: 35489791 Review.
Cited by
-
Microbial Diversity, Community Turnover, and Putative Functions in Submarine Canyon Sediments under the Action of Sedimentary Geology.Microbiol Spectr. 2023 Feb 21;11(2):e0421022. doi: 10.1128/spectrum.04210-22. Online ahead of print. Microbiol Spectr. 2023. PMID: 36802161 Free PMC article.
-
Active Anaerobic Archaeal Methanotrophs in Recently Emerged Cold Seeps of Northern South China Sea.Front Microbiol. 2020 Dec 16;11:612135. doi: 10.3389/fmicb.2020.612135. eCollection 2020. Front Microbiol. 2020. PMID: 33391242 Free PMC article.
-
Potential metabolic and genetic interaction among viruses, methanogen and methanotrophic archaea, and their syntrophic partners.ISME Commun. 2022 Jun 28;2(1):50. doi: 10.1038/s43705-022-00135-2. ISME Commun. 2022. PMID: 37938729 Free PMC article.
-
Niche Differentiation of Sulfate- and Iron-Dependent Anaerobic Methane Oxidation and Methylotrophic Methanogenesis in Deep Sea Methane Seeps.Front Microbiol. 2020 Jul 8;11:1409. doi: 10.3389/fmicb.2020.01409. eCollection 2020. Front Microbiol. 2020. PMID: 32733397 Free PMC article.
-
Higher Abundance of Sediment Methanogens and Methanotrophs Do Not Predict the Atmospheric Methane and Carbon Dioxide Flows in Eutrophic Tropical Freshwater Reservoirs.Front Microbiol. 2021 Mar 17;12:647921. doi: 10.3389/fmicb.2021.647921. eCollection 2021. Front Microbiol. 2021. PMID: 33815337 Free PMC article.
References
-
- Hinrichs K.-U., Boetius A. The anaerobic oxidation of methane: new insights in microbial ecology and biogeochemistry. In: Wefer G., Billett D., Hebbeln D., Jørgensen B. B., Schlüter M., Van Weering T., editors. Ocean Margin Systems. Berlin, Germany: Springer; 2002. pp. 457–477.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

