Promoting Cas9 degradation reduces mosaic mutations in non-human primate embryos
- PMID: 28155910
- PMCID: PMC5290468
- DOI: 10.1038/srep42081
Promoting Cas9 degradation reduces mosaic mutations in non-human primate embryos
Abstract
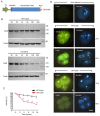
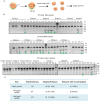
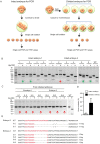
CRISPR-Cas9 is a powerful new tool for genome editing, but this technique creates mosaic mutations that affect the efficiency and precision of its ability to edit the genome. Reducing mosaic mutations is particularly important for gene therapy and precision genome editing. Although the mechanisms underlying the CRSIPR/Cas9-mediated mosaic mutations remain elusive, the prolonged expression and activity of Cas9 in embryos could contribute to mosaicism in DNA mutations. Here we report that tagging Cas9 with ubiquitin-proteasomal degradation signals can facilitate the degradation of Cas9 in non-human primate embryos. Using embryo-splitting approach, we found that shortening the half-life of Cas9 in fertilized zygotes reduces mosaic mutations and increases its ability to modify genomes in non-human primate embryos. Also, injection of modified Cas9 in one-cell embryos leads to live monkeys with the targeted gene modifications. Our findings suggest that modifying Cas9 activity can be an effective strategy to enhance precision genome editing.
Conflict of interest statement
The authors declare no competing financial interests.
Figures


References
-
- Horvath P. & Barrangou R. CRISPR/Cas, the immune system of bacteria and archaea. Science 327, 167–170 (2010). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

