Loss of CD44dim Expression from Early Progenitor Cells Marks T-Cell Lineage Commitment in the Human Thymus
- PMID: 28163708
- PMCID: PMC5247458
- DOI: 10.3389/fimmu.2017.00032
Loss of CD44dim Expression from Early Progenitor Cells Marks T-Cell Lineage Commitment in the Human Thymus
Abstract
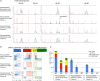
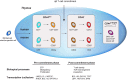
Human T-cell development is less well studied than its murine counterpart due to the lack of genetic tools and the difficulty of obtaining cells and tissues. Here, we report the transcriptional landscape of 11 immature, consecutive human T-cell developmental stages. The changes in gene expression of cultured stem cells on OP9-DL1 match those of ex vivo isolated murine and human thymocytes. These analyses led us to define evolutionary conserved gene signatures that represent pre- and post-αβ T-cell commitment stages. We found that loss of dim expression of CD44 marks human T-cell commitment in early CD7+CD5+CD45dim cells, before the acquisition of CD1a surface expression. The CD44-CD1a- post-committed thymocytes have initiated in frame T-cell receptor rearrangements that are accompanied by loss of capacity to differentiate toward myeloid, B- and NK-lineages, unlike uncommitted CD44dimCD1a- thymocytes. Therefore, loss of CD44 represents a previously unrecognized human thymocyte stage that defines the earliest committed T-cell population in the thymus.
Keywords: CD44; OP9-DL1; T-cell commitment; gene expression; human T-cell development; multi-lineage potential; thymus.
Figures






References
-
- Plum J, De Smedt M, Defresne MP, Leclercq G, Vandekerckhove B. Human CD34+ fetal liver stem cells differentiate to T cells in a mouse thymic microenvironment. Blood (1994) 84(5):1587–93. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

