SeedUSoon: A New Software Program to Improve Seed Stock Management and Plant Line Exchanges between Research Laboratories
- PMID: 28163712
- PMCID: PMC5247430
- DOI: 10.3389/fpls.2017.00013
SeedUSoon: A New Software Program to Improve Seed Stock Management and Plant Line Exchanges between Research Laboratories
Abstract
Plant research is supported by an ever-growing collection of mutant or transgenic lines. In the past, a typical basic research laboratory would focus on only a few plant lines that were carefully isolated from collections of lines containing random mutations. The subsequent technological breakthrough in high-throughput sequencing, combined with novel and highly efficient mutagenesis techniques (including site-directed mutagenesis), has led to a recent exponential growth in plant line collections used by individual researchers. Tracking the generation and genetic properties of these genetic resources is thus becoming increasingly challenging for researchers. Another difficulty for researchers is controlling the use of seeds protected by a Material Transfer Agreement, as often only the original recipient of the seeds is aware of the existence of such documents. This situation can thus lead to difficult legal situations. Simultaneously, various institutions and the general public now demand more information about the use of genetically modified organisms (GMOs). In response, researchers are seeking new database solutions to address the triple challenge of research competition, legal constraints, and institutional/public demands. To help plant biology laboratories organize, describe, store, trace, and distribute their seeds, we have developed the new program SeedUSoon, with simplicity in mind. This software contains data management functions that allow the separate tracking of distinct mutations, even in successive crossings or mutagenesis. SeedUSoon reflects the biotechnological diversity of mutations and transgenes contained in any specific line, and the history of their inheritance. It can facilitate GMO certification procedures by distinguishing mutations on the basis of the presence/absence of a transgene, and by recording the technology used for their generation. Its interface can be customized to match the context and rules of any laboratory. In addition, SeedUSoon includes functions to help the laboratory protect intellectual property, export data, and facilitate seed exchange between laboratories. The SeedUSoon program, which is customizable to match individual practices and preferences, provides a powerful toolkit to plant laboratories searching for innovative approaches in laboratory management.
Keywords: MTA; database; genealogy; genetics; plant; seed; software.
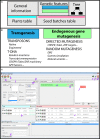
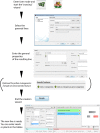
Figures






Similar articles
-
Phytotracker, an information management system for easy recording and tracking of plants, seeds and plasmids.Plant Methods. 2012 Oct 13;8(1):43. doi: 10.1186/1746-4811-8-43. Plant Methods. 2012. PMID: 23062011 Free PMC article.
-
Proceedings of the Second Workshop on Theory meets Industry (Erwin-Schrödinger-Institute (ESI), Vienna, Austria, 12-14 June 2007).J Phys Condens Matter. 2008 Feb 13;20(6):060301. doi: 10.1088/0953-8984/20/06/060301. Epub 2008 Jan 24. J Phys Condens Matter. 2008. PMID: 21693862
-
ScienceLabDatabase: a computer program to organize a molecular biology laboratory.Biotechniques. 1999 Jun;26(6):1186-91. doi: 10.2144/99266bc05. Biotechniques. 1999. PMID: 10376159
-
New ECCO model documents for Material Deposit and Transfer Agreements in compliance with the Nagoya Protocol.FEMS Microbiol Lett. 2020 Mar 1;367(5):fnaa044. doi: 10.1093/femsle/fnaa044. FEMS Microbiol Lett. 2020. PMID: 32149346 Free PMC article. Review.
-
Genetically modified seeds and plant propagating material in Europe: potential routes of entrance and current status.Heliyon. 2019 Feb 15;5(2):e01242. doi: 10.1016/j.heliyon.2019.e01242. eCollection 2019 Feb. Heliyon. 2019. PMID: 30815609 Free PMC article. Review.
Cited by
-
SHiNeMaS: a web tool dedicated to seed lots history, phenotyping and cultural practices.Plant Methods. 2020 Jul 23;16:98. doi: 10.1186/s13007-020-00640-2. eCollection 2020. Plant Methods. 2020. PMID: 32714430 Free PMC article.
-
Sustainable Agriculture through Multidisciplinary Seed Nanopriming: Prospects of Opportunities and Challenges.Cells. 2021 Sep 15;10(9):2428. doi: 10.3390/cells10092428. Cells. 2021. PMID: 34572078 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

