Structural basis of the specificity of USP18 toward ISG15
- PMID: 28165509
- PMCID: PMC5405867
- DOI: 10.1038/nsmb.3371
Structural basis of the specificity of USP18 toward ISG15
Abstract
Protein modification by ubiquitin and ubiquitin-like modifiers (Ubls) is counteracted by ubiquitin proteases and Ubl proteases, collectively termed DUBs. In contrast to other proteases of the ubiquitin-specific protease (USP) family, USP18 shows no reactivity toward ubiquitin but specifically deconjugates the interferon-induced Ubl ISG15. To identify the molecular determinants of this specificity, we solved the crystal structures of mouse USP18 alone and in complex with mouse ISG15. USP18 was crystallized in an open and a closed conformation, thus revealing high flexibility of the enzyme. Structural data, biochemical and mutational analysis showed that only the C-terminal ubiquitin-like domain of ISG15 is recognized and essential for USP18 activity. A critical hydrophobic patch in USP18 interacts with a hydrophobic region unique to ISG15, thus providing evidence that USP18's ISG15 specificity is mediated by a small interaction interface. Our results may provide a structural basis for the development of new drugs modulating ISG15 linkage.
Conflict of interest statement
The authors declare no competing financial interests.
Figures


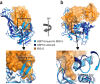
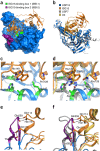


References
-
- Haas AL, Ahrens P, Bright PM, Ankel H. Interferon induces a 15-kilodalton protein exhibiting marked homology to ubiquitin. J. Biol. Chem. 1987;262:11315–11323. - PubMed
-
- Narasimhan J, et al. Crystal structure of the interferon-induced ubiquitin-like protein ISG15. J. Biol. Chem. 2005;280:27356–27365. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

