Phyx: phylogenetic tools for unix
- PMID: 28174903
- PMCID: PMC5870855
- DOI: 10.1093/bioinformatics/btx063
Phyx: phylogenetic tools for unix
Abstract
Summary: The ease with which phylogenomic data can be generated has drastically escalated the computational burden for even routine phylogenetic investigations. To address this, we present phyx : a collection of programs written in C ++ to explore, manipulate, analyze and simulate phylogenetic objects (alignments, trees and MCMC logs). Modelled after Unix/GNU/Linux command line tools, individual programs perform a single task and operate on standard I/O streams that can be piped to quickly and easily form complex analytical pipelines. Because of the stream-centric paradigm, memory requirements are minimized (often only a single tree or sequence in memory at any instance), and hence phyx is capable of efficiently processing very large datasets.
Availability and implementation: phyx runs on POSIX-compliant operating systems. Source code, installation instructions, documentation and example files are freely available under the GNU General Public License at https://github.com/FePhyFoFum/phyx.
Contact: eebsmith@umich.edu.
Supplementary information: Supplementary data are available at Bioinformatics online.
© The Author 2017. Published by Oxford University Press.
Figures
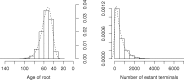
References
-
- Bollback J.P. (2002) Bayesian model adequacy and choice in phylogenetics. Mol. Biol. Evol., 19, 1171–1180. - PubMed
-
- Maddison W.P., Maddison D.R. (2017) Mesquite: a modular system for evolutionary analysis. Version 3.2, http://mesquiteproject.org.
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

