Transcriptional firing helps to drive NETosis
- PMID: 28176807
- PMCID: PMC5296899
- DOI: 10.1038/srep41749
Transcriptional firing helps to drive NETosis
Abstract
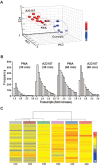
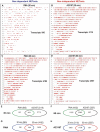
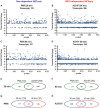
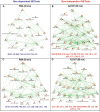
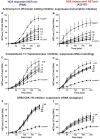
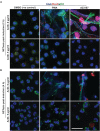
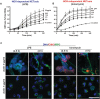
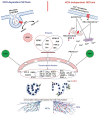
Neutrophils are short-lived innate immune cells. These cells respond quickly to stimuli, and die within minutes to hours; the relevance of DNA transcription in dying neutrophils remains an enigma for several decades. Here we show that the transcriptional activity reflects the degree of DNA decondensation occurring in both NADPH oxidase 2 (Nox)-dependent and Nox-independent neutrophil extracellular trap (NET) formation or NETosis. Transcriptomics analyses show that transcription starts at multiple loci in all chromosomes earlier in the rapid Nox-independent NETosis (induced by calcium ionophore A23187) than Nox-dependent NETosis (induced by PMA). NETosis-specific kinase cascades differentially activate transcription of different sets of genes. Inhibitors of transcription, but not translation, suppress both types of NETosis. In particular, promoter melting step is important to drive NETosis (induced by PMA, E. coli LPS, A23187, Streptomyces conglobatus ionomycin). Extensive citrullination of histones in multiple loci occurs only during calcium-mediated NETosis, suggesting that citrullination of histone contributes to the rapid DNA decondensation seen in Nox-independent NETosis. Furthermore, blocking transcription suppresses both types of NETosis, without affecting the reactive oxygen species production that is necessary for antimicrobial functions. Therefore, we assign a new function for transcription in neutrophils: Transcriptional firing, regulated by NETosis-specific kinases, helps to drive NETosis.
Conflict of interest statement
N.P. has filed a patent to use nucleic acid metabolism inhibitors to inhibit NETosis.
Figures
Similar articles
-
Histone Acetylation Promotes Neutrophil Extracellular Trap Formation.Biomolecules. 2019 Jan 18;9(1):32. doi: 10.3390/biom9010032. Biomolecules. 2019. PMID: 30669408 Free PMC article.
-
Alkaline pH Promotes NADPH Oxidase-Independent Neutrophil Extracellular Trap Formation: A Matter of Mitochondrial Reactive Oxygen Species Generation and Citrullination and Cleavage of Histone.Front Immunol. 2018 Jan 9;8:1849. doi: 10.3389/fimmu.2017.01849. eCollection 2017. Front Immunol. 2018. PMID: 29375550 Free PMC article.
-
Furanoid F-Acid F6 Uniquely Induces NETosis Compared to C16 and C18 Fatty Acids in Human Neutrophils.Biomolecules. 2018 Nov 13;8(4):144. doi: 10.3390/biom8040144. Biomolecules. 2018. PMID: 30428625 Free PMC article.
-
How Do ROS Induce NETosis? Oxidative DNA Damage, DNA Repair, and Chromatin Decondensation.Biomolecules. 2024 Oct 16;14(10):1307. doi: 10.3390/biom14101307. Biomolecules. 2024. PMID: 39456240 Free PMC article. Review.
-
Post-Translational Modifications in NETosis and NETs-Mediated Diseases.Biomolecules. 2019 Aug 14;9(8):369. doi: 10.3390/biom9080369. Biomolecules. 2019. PMID: 31416265 Free PMC article. Review.
Cited by
-
Progression of Cystic Fibrosis Lung Disease from Childhood to Adulthood: Neutrophils, Neutrophil Extracellular Trap (NET) Formation, and NET Degradation.Genes (Basel). 2019 Feb 26;10(3):183. doi: 10.3390/genes10030183. Genes (Basel). 2019. PMID: 30813645 Free PMC article. Review.
-
Hypertonic Saline Suppresses NADPH Oxidase-Dependent Neutrophil Extracellular Trap Formation and Promotes Apoptosis.Front Immunol. 2018 Mar 8;9:359. doi: 10.3389/fimmu.2018.00359. eCollection 2018. Front Immunol. 2018. PMID: 29593709 Free PMC article.
-
Neutrophil Extracellular Trapping Role in Cancer, Metastases, and Cancer-Related Thrombosis: a Narrative Review of the Current Evidence Base.Curr Oncol Rep. 2021 Aug 3;23(10):118. doi: 10.1007/s11912-021-01103-0. Curr Oncol Rep. 2021. PMID: 34342735 Free PMC article. Review.
-
Extracellular DNA traps in a ctenophore demonstrate immune cell behaviors in a non-bilaterian.Nat Commun. 2024 Apr 6;15(1):2990. doi: 10.1038/s41467-024-46807-6. Nat Commun. 2024. PMID: 38582801 Free PMC article.
-
Bio-impedance measurement allows displaying the early stages of neutrophil extracellular traps.EXCLI J. 2020 Nov 5;19:1481-1495. doi: 10.17179/excli2020-2868. eCollection 2020. EXCLI J. 2020. PMID: 33250682 Free PMC article.
References
-
- Takei H., Araki A., Watanabe H., Ichinose A. & Sendo F. Rapid killing of human neutrophils by the potent activator phorbol 12-myristate 13-acetate (PMA) accompanied by changes different from typical apoptosis or necrosis. J Leukoc Biol 59, 229–240 (1996). - PubMed
-
- Brinkmann V. et al.. Neutrophil extracellular traps kill bacteria. Science 303, 1532–1535 (2004). - PubMed
-
- Steinberg B. E. & Grinstein S. Unconventional roles of the NADPH oxidase: signaling, ion homeostasis, and cell death. Sci STKE 2007, pe11 (2007). - PubMed
-
- Hakkim A. et al.. Activation of the Raf-MEK-ERK pathway is required for neutrophil extracellular trap formation. Nat Chem Biol 7, 75–77 (2010). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

