Immortalized human myotonic dystrophy muscle cell lines to assess therapeutic compounds
- PMID: 28188264
- PMCID: PMC5399563
- DOI: 10.1242/dmm.027367
Immortalized human myotonic dystrophy muscle cell lines to assess therapeutic compounds
Abstract
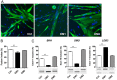
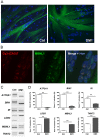
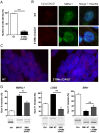
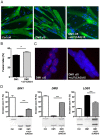
Myotonic dystrophy type 1 (DM1) and type 2 (DM2) are autosomal dominant neuromuscular diseases caused by microsatellite expansions and belong to the family of RNA-dominant disorders. Availability of cellular models in which the DM mutation is expressed within its natural context is essential to facilitate efforts to identify new therapeutic compounds. Here, we generated immortalized DM1 and DM2 human muscle cell lines that display nuclear RNA aggregates of expanded repeats, a hallmark of myotonic dystrophy. Selected clones of DM1 and DM2 immortalized myoblasts behave as parental primary myoblasts with a reduced fusion capacity of immortalized DM1 myoblasts when compared with control and DM2 cells. Alternative splicing defects were observed in differentiated DM1 muscle cell lines, but not in DM2 lines. Splicing alterations did not result from differentiation delay because similar changes were found in immortalized DM1 transdifferentiated fibroblasts in which myogenic differentiation has been forced by overexpression of MYOD1. As a proof-of-concept, we show that antisense approaches alleviate disease-associated defects, and an RNA-seq analysis confirmed that the vast majority of mis-spliced events in immortalized DM1 muscle cells were affected by antisense treatment, with half of them significantly rescued in treated DM1 cells. Immortalized DM1 muscle cell lines displaying characteristic disease-associated molecular features such as nuclear RNA aggregates and splicing defects can be used as robust readouts for the screening of therapeutic compounds. Therefore, immortalized DM1 and DM2 muscle cell lines represent new models and tools to investigate molecular pathophysiological mechanisms and evaluate the in vitro effects of compounds on RNA toxicity associated with myotonic dystrophy mutations.
Keywords: Alternative splicing; Expanded repeats; Muscle cell line; Myotonic dystrophy; Nuclear aggregates; Therapeutic compounds.
© 2017. Published by The Company of Biologists Ltd.
Conflict of interest statement
The authors declare no competing or financial interests.
Figures

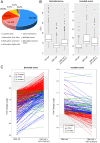
References
-
- Bigot A., Klein A. F., Gasnier E., Jacquemin V., Ravassard P., Butler-Browne G., Mouly V. and Furling D. (2009). Large CTG repeats trigger p16-dependent premature senescence in myotonic dystrophy type 1 muscle precursor cells. Am. J. Pathol. 174, 1435-1442. 10.2353/ajpath.2009.080560 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

