Shoot Removal Induces Chloroplast Development in Roots via Cytokinin Signaling
- PMID: 28193764
- PMCID: PMC5373043
- DOI: 10.1104/pp.16.01368
Shoot Removal Induces Chloroplast Development in Roots via Cytokinin Signaling
Abstract
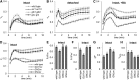
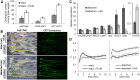
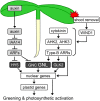
The development of plant chloroplasts is regulated by various developmental, environmental, and hormonal cues. In Arabidopsis (Arabidopsis thaliana), chloroplast development is repressed in roots via auxin signaling. However, roots develop chloroplasts when they are detached from the shoot. In contrast to auxin, cytokinin positively affects chloroplast development in roots, but the role and signaling pathway of cytokinin in the root greening response remain unclear. To understand the regulatory pathways of chloroplast development in the plant stress response, we examined the mechanisms underlying the conditional greening of detached roots. In wild-type Arabidopsis roots, shoot removal activates type B ARABIDOPSIS RESPONSE REGULATOR (ARR)-mediated cytokinin signaling and induces chlorophyll accumulation and photosynthetic remodeling. ARR1 and ARR12 are essential for up-regulating nucleus- and plastid-encoded genes associated with chloroplast development in detached roots. In this process, WOUND INDUCED DEDIFFERENTIATION1 and class B GATA transcription factors (B-GATAs) act upstream and downstream of ARRs, respectively. Overexpression of B-GATAs promotes root greening, as does shoot removal, dependent on a light signaling transcription factor, LONG HYPOCOTYL5. Auxin represses the root greening response independent of ARR signaling. GNC-LIKE (GNL), a B-GATA, is strongly up-regulated in detached roots via ARR1 and ARR12 but is repressed by auxin, so GNL may function at the point of convergence of cytokinin and auxin signaling in the root greening response.
© 2017 American Society of Plant Biologists. All Rights Reserved.
Figures
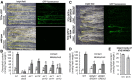
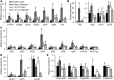
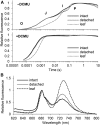

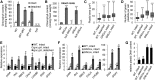
Similar articles
-
Cytokinin Regulates the Etioplast-Chloroplast Transition through the Two-Component Signaling System and Activation of Chloroplast-Related Genes.Plant Physiol. 2016 Sep;172(1):464-78. doi: 10.1104/pp.16.00640. Epub 2016 Jul 7. Plant Physiol. 2016. PMID: 27388681 Free PMC article.
-
Regulation of root greening by light and auxin/cytokinin signaling in Arabidopsis.Plant Cell. 2012 Mar;24(3):1081-95. doi: 10.1105/tpc.111.092254. Epub 2012 Mar 13. Plant Cell. 2012. PMID: 22415275 Free PMC article.
-
Functional characterization of type-B response regulators in the Arabidopsis cytokinin response.Plant Physiol. 2013 May;162(1):212-24. doi: 10.1104/pp.112.208736. Epub 2013 Mar 12. Plant Physiol. 2013. PMID: 23482873 Free PMC article.
-
Regulation of chloroplast development and function by cytokinin.J Exp Bot. 2015 Aug;66(16):4999-5013. doi: 10.1093/jxb/erv132. Epub 2015 Apr 4. J Exp Bot. 2015. PMID: 25873684 Review.
-
Arabidopsis cytokinin signaling pathway.Sci STKE. 2007 Oct 9;2007(407):cm5. doi: 10.1126/stke.4072007cm5. Sci STKE. 2007. PMID: 17925576 Review.
Cited by
-
Static magnetic field regulates Arabidopsis root growth via auxin signaling.Sci Rep. 2019 Oct 7;9(1):14384. doi: 10.1038/s41598-019-50970-y. Sci Rep. 2019. PMID: 31591431 Free PMC article.
-
Cytosolic GLUTAMINE SYNTHETASE1;1 Modulates Metabolism and Chloroplast Development in Roots.Plant Physiol. 2020 Apr;182(4):1894-1909. doi: 10.1104/pp.19.01118. Epub 2020 Feb 5. Plant Physiol. 2020. PMID: 32024696 Free PMC article.
-
Albino lethal 13, a chloroplast-imported protein required for chloroplast development in rice.Plant Direct. 2024 Jun 20;8(6):e610. doi: 10.1002/pld3.610. eCollection 2024 Jun. Plant Direct. 2024. PMID: 38903415 Free PMC article.
-
An Artificial Conversion of Roots into Organs with Shoot Stem Characteristics by Inducing Two Transcription Factors.iScience. 2020 Jul 24;23(7):101332. doi: 10.1016/j.isci.2020.101332. Epub 2020 Jul 14. iScience. 2020. PMID: 32668199 Free PMC article.
-
Fine Mapping and Transcriptome Analysis of Virescent Leaf Gene v-2 in Cucumber (Cucumis sativus L.).Front Plant Sci. 2020 Sep 25;11:570817. doi: 10.3389/fpls.2020.570817. eCollection 2020. Front Plant Sci. 2020. PMID: 33101337 Free PMC article.
References
-
- Chiang YH, Zubo YO, Tapken W, Kim HJ, Lavanway AM, Howard L, Pilon M, Kieber JJ, Schaller GE (2012) Functional characterization of the GATA transcription factors GNC and CGA1 reveals their key role in chloroplast development, growth, and division in Arabidopsis. Plant Physiol 160: 332–348 - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

