Genome Editing in hPSCs Reveals GATA6 Haploinsufficiency and a Genetic Interaction with GATA4 in Human Pancreatic Development
- PMID: 28196600
- PMCID: PMC5419850
- DOI: 10.1016/j.stem.2017.01.001
Genome Editing in hPSCs Reveals GATA6 Haploinsufficiency and a Genetic Interaction with GATA4 in Human Pancreatic Development
Abstract
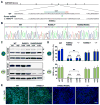
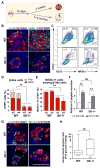
Human disease phenotypes associated with haploinsufficient gene requirements are often not recapitulated well in animal models. Here, we have investigated the association between human GATA6 haploinsufficiency and a wide range of clinical phenotypes that include neonatal and adult-onset diabetes using CRISPR (clustered regularly interspaced short palindromic repeat)/Cas9-mediated genome editing coupled with human pluripotent stem cell (hPSC) directed differentiation. We found that loss of one GATA6 allele specifically affects the differentiation of human pancreatic progenitors from the early PDX1+ stage to the more mature PDX1+NKX6.1+ stage, leading to impaired formation of glucose-responsive β-like cells. In addition to this GATA6 haploinsufficiency, we also identified dosage-sensitive requirements for GATA6 and GATA4 in the formation of both definitive endoderm and pancreatic progenitor cells. Our work expands the application of hPSCs from studying the impact of individual gene loci to investigation of multigenic human traits, and it establishes an approach for identifying genetic modifiers of human disease.
Keywords: CRISPR/Cas9 genome editing; GATA6 and GATA4; definitive endoderm; genetic modifier; haploinsufficiency; human embryonic stem cells; human pluripotent stem cells disease modeling; insulin producing pancreatic beta cells; pancreatic agenesis and neonatal diabetes; pancreatic progenitor.
Copyright © 2017 Elsevier Inc. All rights reserved.
Figures




Comment in
-
Gotta Have GATA for Human Pancreas Development.Cell Stem Cell. 2017 May 4;20(5):577-579. doi: 10.1016/j.stem.2017.04.004. Cell Stem Cell. 2017. PMID: 28475878
Similar articles
-
GATA6 Cooperates with EOMES/SMAD2/3 to Deploy the Gene Regulatory Network Governing Human Definitive Endoderm and Pancreas Formation.Stem Cell Reports. 2019 Jan 8;12(1):57-70. doi: 10.1016/j.stemcr.2018.12.003. Stem Cell Reports. 2019. PMID: 30629940 Free PMC article.
-
Gotta Have GATA for Human Pancreas Development.Cell Stem Cell. 2017 May 4;20(5):577-579. doi: 10.1016/j.stem.2017.04.004. Cell Stem Cell. 2017. PMID: 28475878
-
Pancreas-specific deletion of mouse Gata4 and Gata6 causes pancreatic agenesis.J Clin Invest. 2012 Oct;122(10):3516-28. doi: 10.1172/JCI63352. Epub 2012 Sep 24. J Clin Invest. 2012. PMID: 23006325 Free PMC article.
-
CRISPR/Cas9 genome editing in human pluripotent stem cells: Harnessing human genetics in a dish.Dev Dyn. 2016 Jul;245(7):788-806. doi: 10.1002/dvdy.24414. Epub 2016 Jun 9. Dev Dyn. 2016. PMID: 27145095 Review.
-
Human pluripotent stem cell based islet models for diabetes research.Best Pract Res Clin Endocrinol Metab. 2015 Dec;29(6):899-909. doi: 10.1016/j.beem.2015.10.012. Epub 2015 Oct 30. Best Pract Res Clin Endocrinol Metab. 2015. PMID: 26696518 Review.
Cited by
-
Regulation of multiple signaling pathways promotes the consistent expansion of human pancreatic progenitors in defined conditions.Elife. 2024 Jan 5;12:RP89962. doi: 10.7554/eLife.89962. Elife. 2024. PMID: 38180318 Free PMC article.
-
GATA6 Plays an Important Role in the Induction of Human Definitive Endoderm, Development of the Pancreas, and Functionality of Pancreatic β Cells.Stem Cell Reports. 2017 Mar 14;8(3):589-604. doi: 10.1016/j.stemcr.2016.12.026. Epub 2017 Feb 9. Stem Cell Reports. 2017. PMID: 28196690 Free PMC article.
-
GATA6 regulates WNT and BMP programs to pattern precardiac mesoderm during the earliest stages of human cardiogenesis.Elife. 2025 Mar 13;13:RP100797. doi: 10.7554/eLife.100797. Elife. 2025. PMID: 40080060 Free PMC article.
-
GATA6 defines endoderm fate by controlling chromatin accessibility during differentiation of human-induced pluripotent stem cells.Cell Rep. 2021 May 18;35(7):109145. doi: 10.1016/j.celrep.2021.109145. Cell Rep. 2021. PMID: 34010638 Free PMC article.
-
In vitro beta-cell killing models using immune cells and human pluripotent stem cell-derived islets: Challenges and opportunities.Front Endocrinol (Lausanne). 2023 Jan 16;13:1076683. doi: 10.3389/fendo.2022.1076683. eCollection 2022. Front Endocrinol (Lausanne). 2023. PMID: 36726462 Free PMC article. Review.
References
-
- Bonnefond A, Sand O, Guerin B, Durand E, De Graeve F, Huyvaert M, Rachdi L, Kerr-Conte J, Pattou F, Vaxillaire M, et al. GATA6 inactivating mutations are associated with heart defects and, inconsistently, with pancreatic agenesis and diabetes. Diabetologia. 2012;55:2845–2847. - PubMed
-
- Bruneau BG. The developmental genetics of congenital heart disease. Nature. 2008;451:943–948. - PubMed
-
- Bui PH, Dorrani N, Wong D, Perens G, Dipple KM, Quintero-Rivera F. First report of a de novo 18q11.2 microdeletion including GATA6 associated with complex congenital heart disease and renal abnormalities. Am J Med Genet A. 2013;161A:1773–1778. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

