Unusual C΄/D΄ motifs enable box C/D snoRNPs to modify multiple sites in the same rRNA target region
- PMID: 28204564
- PMCID: PMC5389607
- DOI: 10.1093/nar/gkw842
Unusual C΄/D΄ motifs enable box C/D snoRNPs to modify multiple sites in the same rRNA target region
Abstract
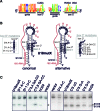
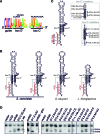
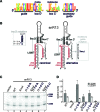
Eukaryotic box C/D small nucleolar (sno)RNPs catalyse the site-specific 2΄-O-methylation of ribosomal RNA. The RNA component (snoRNA) contains guide regions that base-pair with the target site to select the single nucleotide to be modified. The terminal C/D and internal C΄/D΄ motifs in the snoRNA, adjacent to the guide region, function as binding sites for the snoRNP proteins including the enzymatic subunit fibrillarin/Nop1. Four yeast snoRNAs are unusual in that they are predicted to methylate two nucleotides in a single target region. In each case, the internal C΄/D΄ motifs from these snoRNAs differ from the consensus. Our data indicate that the C΄/D΄ motifs in snR13, snR48 and U18 form two alternative structures that lead to differences in the position of the proteins bound to this motif. We propose that each snoRNA forms two different snoRNPs, subtly different in how the proteins are bound to the C΄/D΄ motif, leading to 2΄-O-methylation of different nucleotides in the target region. For snR48 and U18, the unusual C΄/D΄ alone is enough for the modification of two nucleotides. However, for the snR13 snoRNA the unusual C΄/D΄ motif and extra base-pairing, which stimulates rRNA 2΄-O-methylation, are both critical for multiple modifications in the target region.
Figures



Similar articles
-
Box C/D snoRNP catalysed methylation is aided by additional pre-rRNA base-pairing.EMBO J. 2011 May 10;30(12):2420-30. doi: 10.1038/emboj.2011.148. EMBO J. 2011. PMID: 21556049 Free PMC article.
-
AtNUFIP, an essential protein for plant development, reveals the impact of snoRNA gene organisation on the assembly of snoRNPs and rRNA methylation in Arabidopsis thaliana.Plant J. 2011 Mar;65(5):807-19. doi: 10.1111/j.1365-313X.2010.04468.x. Epub 2011 Jan 24. Plant J. 2011. PMID: 21261762
-
The box C/D and H/ACA snoRNPs: key players in the modification, processing and the dynamic folding of ribosomal RNA.Wiley Interdiscip Rev RNA. 2012 May-Jun;3(3):397-414. doi: 10.1002/wrna.117. Epub 2011 Nov 7. Wiley Interdiscip Rev RNA. 2012. PMID: 22065625 Review.
-
Archaeal homologs of eukaryotic methylation guide small nucleolar RNAs: lessons from the Pyrococcus genomes.J Mol Biol. 2000 Apr 7;297(4):895-906. doi: 10.1006/jmbi.2000.3593. J Mol Biol. 2000. PMID: 10736225
-
Implications of small nucleolar RNA-protein complexes discoveries.Recent Pat DNA Gene Seq. 2008;2(1):1-5. doi: 10.2174/187221508783406576. Recent Pat DNA Gene Seq. 2008. PMID: 19075938 Review.
Cited by
-
The large repertoire of 2'-O-methylation guided by C/D snoRNAs on Trypanosoma brucei rRNA.RNA Biol. 2020 Jul;17(7):1018-1039. doi: 10.1080/15476286.2020.1750842. Epub 2020 Apr 21. RNA Biol. 2020. PMID: 32250712 Free PMC article.
-
SnoRNA guide activities: real and ambiguous.RNA. 2021 Nov;27(11):1363-1373. doi: 10.1261/rna.078916.121. Epub 2021 Aug 12. RNA. 2021. PMID: 34385348 Free PMC article.
-
Tuning the ribosome: The influence of rRNA modification on eukaryotic ribosome biogenesis and function.RNA Biol. 2017 Sep 2;14(9):1138-1152. doi: 10.1080/15476286.2016.1259781. Epub 2016 Dec 2. RNA Biol. 2017. PMID: 27911188 Free PMC article. Review.
-
Profiling of RNA ribose methylation in Arabidopsis thaliana.Nucleic Acids Res. 2021 Apr 19;49(7):4104-4119. doi: 10.1093/nar/gkab196. Nucleic Acids Res. 2021. PMID: 33784398 Free PMC article.
-
Biology and clinical relevance of noncoding sno/scaRNAs.Trends Cardiovasc Med. 2018 Feb;28(2):81-90. doi: 10.1016/j.tcm.2017.08.002. Epub 2017 Aug 12. Trends Cardiovasc Med. 2018. PMID: 28869095 Free PMC article. Review.
References
-
- Decatur W.A., Fournier M.J.. rRNA modifications and ribosome function. Trends Biochem. Sci. 2002; 27:344–351. - PubMed
-
- Fischer N., Neumann P., Konevega A.L., Bock L.V., Ficner R., Rodnina M.V., Stark H.. Structure of the E. coli ribosome-EF-Tu complex at <3 A resolution by C-corrected cryo-EM. Nature. 2015; 520:567–570. - PubMed
-
- Motorin Y., Helm M.. RNA nucleotide methylation. Wiley Interdiscip. Rev. RNA. 2011; 2:611–631. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

